Keiichi Ito1*, Kenichi Hongo1, Taro Date1, Satoshi Morimoto1, Akira Yoshii1, Haruka Kimura1, Yoshiro Tanaka1, Tomohisa Nagoshi1, Yoichiro Kusakari2, Toru Akaike2, Susumu Minamisawa2, Kazuo Ogawa1, Kosuke Minai1, Makoto Kawai1, Seiichiro Matsuo1, Teiichi Yamane1and Michihiro Yoshimura1
1Division of Cardiology, Department of Internal Medicine, The Jikei University School of Medicine, Nishi-shinbashi, Minato-ku, Tokyo, Japan
2Department of Cell Physiology, The Jikei University School of Medicine, Nishi-shinbashi, Minato-ku, Tokyo, Japan
*Corresponding Author:
Keiichi Ito
Division of Cardiology, Department of Internal Medicine, The Jikei University School of Medicine, 3-25-8, Nishi-shinbashi, Minato-ku, Tokyo 105-8461, Japan
Tel: +81-3-3433-1111
Fax: +81-3-3459-6043
E-mail: keke-ito@jikei.ac.jp
Rec date: May 15, 2018; Acc date: May 25, 2018; Pub date: May 28, 2018
Citation: Ito K, Hongo K, Date T, Morimoto S, Yoshii A, et al. (2018) Mice with the Heart-Specific Overexpression of Prothrombin-Do Not Show Cardiac Insufficiency. Transl Biomed. Vol.9 No.2.148. doi: 10.21767/2172-0479.100148.
Letter to Editor
We previously reported that thrombin exists in the heart [1]. We also reported that thrombin is involved in the pathogenesis of dilated cardiomyopathy (DCM) [2]. We previously investigated the effects of dabigatran (a direct thrombin inhibitor) and found that it has a protective effect against the pathogenesis of dilated cardiomyopathy. Thus, we next focused on the precursor protein of thrombin, prothrombin. Prothrombin is universally distributed in various tissues, including heart tissue [3]. We therefore attempted to generate a mouse model of heart-specific prothrombin overexpression and examined the phenotype of those animals. All the animals received humane care. The study protocols complied with the guidelines of the Animal Research Committee at The Jikei University School of Medicine (26-021). The present study was supported by JSPS KAKENHI Grant Number 26860589, the Uehara Memorial Foundation and The Jikei University Research Fund.
We generated a mouse model of heart-specific prothrombin overexpression (PT-TG mice). The cDNA encoding mouse prothrombin was subcloned into the αMHC (myosin heavy chain) expression cassette via Xba/EcoRsites and transgenic mice (C57BL/6) were generated and screened by Southern blotting of DNA from tail biopsy specimens using two primer mixtures. We used 8-week-old PT-TG mice and wild-type mice. 4-week-old and 8-week-old mice were subjected to transthoracic echocardiography using a Vivid7 echocardiogram unit (General Electric, Tampa, FL). Systolic blood pressure and heart rate of 8-week old wild-type mice were monitored with a tail-cuff system (BP-98A, Softron, Tokyo, Japan). The real-time PCR protocol consisted of one cycle at 95°C for 20 s followed by 40 cycles at 95°C for 1 s and 60°C for 20 s using primers for prothrombin (Applied Biosystems, Mm00438843_m1) and Glyceraldehyde 3- phosphate dehydrogenase (GAPDH) (Applied Biosystems, Mm03302249_g1). The transcriptional levels were determined using the ΔΔCt method, with normalization to the transcriptional level of GAPDH. The results are presented as the mean ± standard error of the mean, and comparisons between two groups were made using the Wilcoxon test. Survival curves were generated using the Kaplan-Meier method (MedCalc Software, Mariakerke, Belgium). P values of <0.05 were considered to indicate statistical significance. All of the statistical analyses (other than the survival curves) were performed using the SPSS software program (version 21, SPSS Japan Inc., Tokyo, Japan).
The expression levels of prothrombin in the PT-TG mice significantly increased in comparison to the wild-type mice (Figure 1A).
We examined the prothrombin mRNA in the wild-type mice, however, we could not detect prothrombin mRNA in the heart (Figure 1A). We further analysed the liver prothrombin mRNA in the mice of the wild-type group as positive controls and detected a large amount of prothrombin mRNA (Figure 1B). There were no significant differences in the heart weights between the wild-type mice and the PT-TG mice (Table 1). The echocardiographic investigations showed no significant differences in fractional shortening or in the diameter of the left ventricle (Table 1). Systolic blood pressure and heart rate of 8-week old wild-type mice were 105.3 ± 3.3 mmHg and 611.5 ± 22.4 beats/min, respectively, which means the procedure is not causing changes that could influence the tests (n=6). The heart-specific overexpression of prothrombin did not worsen their life span and no thromboembolic complications were observed (Figure 1C).
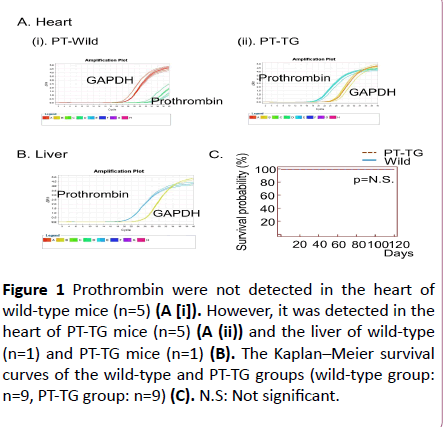
Figure 1: Prothrombin were not detected in the heart of wild-type mice (n=5) (A [i]). However, it was detected in the heart of PT-TG mice (n=5) (A (ii)) and the liver of wild-type (n=1) and PT-TG mice (n=1) (B). The Kaplan–Meier survival curves of the wild-type and PT-TG groups (wild-type group: n=9, PT-TG group: n=9) (C). N.S: Not significant.
| Comparison |
Wild |
PT-TG |
| BW (g) |
18.1 ± 1.1 |
17.0 ± 1.7 |
| HW (mg) |
101.4 ± 10.6 |
89.9 ± 16.6 |
| HW/BW (X10-3) |
5.6 ± 0.4 |
5.3 ± 0.5 |
| Echocardiographic data |
| IVSd (mm) |
0.67 ± 0.21 |
0.69 ± 0.21 |
| PWd (mm) |
0.67 ± 0.24 |
0.70 ± 0.22 |
| LVDd (mm) |
31.2 ± 1.0 |
32.0 ± 1.1 |
| LVDs (mm) |
16.5 ± 0.8 |
17.1 ± 0.8 |
| FS (%) |
47.1 ± 1.2 |
46.7 ± 0.9 |
BW: Body weight; HW: Heart Weight; IVSd: Diastolic, septal wall thicknes; PWd: Diastolic posterior wall thickness; LVDd: Left Ventricular Diastolic Dimension; LVDs: Left Ventricular Systolic Dimension; FS: Fractional Shortening; PT-TG: Prothrombin Transgenic. Body weight and heart weight (Wild group: n=5, PT-TG group: n=5) and echocardiographic data (Wild group: n=4, PT-TG group: n=4) were investigated.
Table 1: Comparison of body weight, heart weight and functional indices of echocardiography between Wild group and PT-TG group.
We previously reported that tissue thrombin exists in the normal human heart by immunohistochemical analysis [1]. We have also reported that thrombin is involved in the pathology of dilated cardiomyopathy [2]. However, the role of prothrombin which is a precursor of thrombin in the heart remains unclear. Prothrombin has been observed in various tissues, including heart tissue [3]. The congenital knockout of prothrombin was revealed to be lethal in prothrombin knockout mouse model [4,5]. Moreover, in mice, the conditional knockout of prothrombin resulted in severe haemorrhagic events within the heart tissue [6]. In our study, the mRNA of prothrombin was detected in the liver as a positive control. Surprisingly, we could not detect the mRNA of prothrombin in the heart tissue of wild-type mice. On the other hand, abundant prothrombin mRNA was found in the heart tissue of PT-TG mice which implied the success of the mouse model of heart-specific prothrombin overexpression. However, the cardiac function did not change, in spite of the excessive level of prothrombin. Our present observations, however, suggested that rather than being the sole factor in the pathogenesis of DCM, thrombin may be an exacerbating factor. More pathophysiological roles of prothrombin remain to be elucidated.
22700
References
- Ito K, Date T, Ikegami M, Hongo K, Fujisaki M, et al. (2013) An immunohistochemical analysis of tissue thrombin expression in the human atria. PLoS One 8: e65817
- Ito K, Hongo K, Date T, Ikegami M, Hano H, et al. (2017) Tissue thrombin is associated with the pathogenesis of dilated cardiomyopathy. Int J Cardiol 228: 821-827.
- McBane RD, Miller RS, Hassinger NL, Chesebro JH, Nemerson Y, et al. (1997) Tissue prothrombin. Universal distribution in smooth muscle. Arterioscler Thromb Vasc Biol 17: 2430-2436.
- Sun WY, Witte DP, Degen JL, Colbert MC, Burkart MC, et al. (1998) Prothrombin deficiency results in embryonic and neonatal lethality in mice. Proc Natl Acad Sci USA 95: 7597-7602.
- Xue J, Wu Q, Westfield LA, Tuley EA, Lu D, et al. (1998) Incomplete embryonic lethality and fatal neonatal haemorrhage caused by prothrombin deficiency in mice. Proc Natl Acad Sci USA 95: 7603-7607.
- Mullins ES, Kombrinck KW, Talmage KE, Shaw MA, Witte DP, et al. (2009) Genetic elimination of prothrombin in adult mice is not compatible with survival and results in spontaneous haemorrhagic events in both heart and brain. Blood 113: 696-704.