Keywords
Phylogenetic tree; Sequence similarities; Antiviral resistance; DNA mutation
Abbreviations
FJ: Fujian; VN: Vietnam; GX: Guangxi; JX: Jiangxi; HN: Hunan; AKH: Astrakhan; YN: Yunnan; GY: Guiyang; XZ: Xinjiang; KH: Cambodia; EGP: Egypt; NG: Nigeria; BVR: Bavaria; QH: Qinghai; SE: Sweden; LAO: Laos; KW: Kuwait; CK: Chicken; DK: Duck; GS: Goose; SN: Swan; BHGS: Bar-Headed Goose; MDK: Muscovy Duck; MALL: Mallard; WDK: Wild Duck; SN: Swan; MSN: Mute Swan; CGO: Cygnus Olor; TDK: Tufted Duck; GDK: Goldeneye Duck; HGL: Herring Gull
Introduction
Neuraminidase; also referred to as sialidase; is an antigenic glycoprotein anchored in the surface envelope of the influenza virions; which hydrolytically cleaves the terminal sialic acid from the host cell receptors. Thus; it plays a crucial role in the release of viral progeny from the membranes of infected cells; prevents self-aggregation of virions and facilitates the movement of the infectious viral particles in the mucus of the respiratory epithelia [1,2]. Influenza neuraminidase has been established as a key drug target for the prophylaxis and treatment of influenza infections; predominantly for the following reasons: Firstly; the structure of the influenza neuraminidase active site is highly conserved between influenza A and B strains; making neuraminidase an attractive target for the development of broad-spectrum inhibitors [3]. Secondly; resistance to neuraminidase inhibitors develops less commonly than to other anti-influenza drugs. Nevertheless; the intensive application of neuraminidase inhibitors for influenza treatment results in a permanently increasing number of drug-resistant strains [4]. Thirdly; in contrast to amantadines; neuraminidase inhibitors are mostly well tolerated in patients under therapy [5]. Finally; neuraminidase protein is a freely accessible target for antiviral molecules with an extracellular mode of action.
The anti-influenza drugs are usually classified according to their target in the viral life-cycle. Such antiviral molecules are particularly used as inhibitors of the following processes: attachment of the virus to host cell receptors; endocytosis and fusion of viral and cell membranes; replication and transcription of the viral genome; synthesis of viral proteins; assembly of the viral progeny and release of the new virions into the outside environment. The following paragraphs are focused on the description of basic classes of influenza virus inhibitors.
Avian influenza A consists of two major glycoproteins which are Hemaglutinin (HA) and Neuraminidase (NA) [6]. HA glycoproteins are more prone to attach to the cell surface sialic acid receptors. There is a difference between host surface receptors on the target cell which is believed to be the possible restrictive factor of avian influenza. HA gene of avian cell binds to Sia2-3 Galactose containing receptor which is different from human Sia2- 6Galactose containing receptor [7]. Before functioning as a virus it needs post translational cleavage by host proteases [8]. HA followed by NA are important antigenic determinant from which neutralizing antibodies are directed. There are several subtypes of HA and NA. 18 different HA subtypes (H1 to H18) and 11 different NA subtypes (N1 to N11) are found [9]. There is a membrane protein named M2 protein which regulates the internal PH level of the virus. This membrane protein is responsible for uncoating the virus during early stages of viral replication [10]. Amantadine and rimantadine block this function. NA catalyze the cleavage of glycosidic linkages to sialic acid on the surface of the viral particle and host cell thus preventing the aggregation and facilitating the release of progeny viruses from the infected cell. Antiviral drugs like Oseltamivir and zanamivir (NA inhibitors) inhibits this important function are the key to the antiviral treatment.
Neuraminidase inhibitors (NAIs) are first-line agents for the treatment and prevention of influenza virus infections. Resistance to the NAIs can be both drug and virus type or subtype specific. The neuraminidase (NA) inhibitors (orally administered oseltamivir and inhaled zanamivir) are currently an important class of antiviral drugs available for the treatment of seasonal and pandemic influenza. Although administration of NA inhibitors may significantly reduce influenza virus transmission; it risks the emergence of drug-resistant variants [11,12]. Several mutations in seasonal or pandemic strains confer resistance to oseltamivir; which is currently the most widely used drug. These mutations are found mainly in the framework of the enzyme and they appear to destabilize drug binding to the target enzyme; thereby reducing viral susceptibility to the treatment [13-15]. Oseltamivir-resistant H5N1 viruses with the H274Y NA mutation have been isolated from three patients during drug treatment or prophylaxis [16,17] and those with the N294S NA mutation; from two patients in Egypt [18]. In addition; the highlypathogenic A⁄ Hanoi ⁄ 30408 ⁄ 2005(H5N1) influenza virus that was isolated from a patient treated with oseltamivir had a mixed oseltamivir-sensitive and oseltamivir-resistant population [19].
Materials and methods: Sequence and data source
Data used in this study are obtained using nucleotide BLAST search from publicly available database of National Centre for Biotechnology Information (NCBI). Multiple sequence alignments; editing; assembly of strains were performed in windows platform with the Geneious program version 7.1.3 (trial). Numbers at nodes in the tree indicate Neighbor-Joining bootstraps value generated from 1,000 replicates.
Result and Discussion
In 2005 we have selected total 169 strains of Neuraminidase (NA) strain of H5N1 (Figure 1 and Table 1).
| Accession number |
Strain name |
| EU930918 |
DK/VN/205/05 |
| EF124227 |
GS/GX/5414/05 |
| HM172200 |
DK/JX/80/05 |
| EU329179 |
WDK/HN/021/05 |
| EF124280 |
GS/GX/3316/05 |
Table 1: List of diverse strains from our analysis of 2005 are given below.
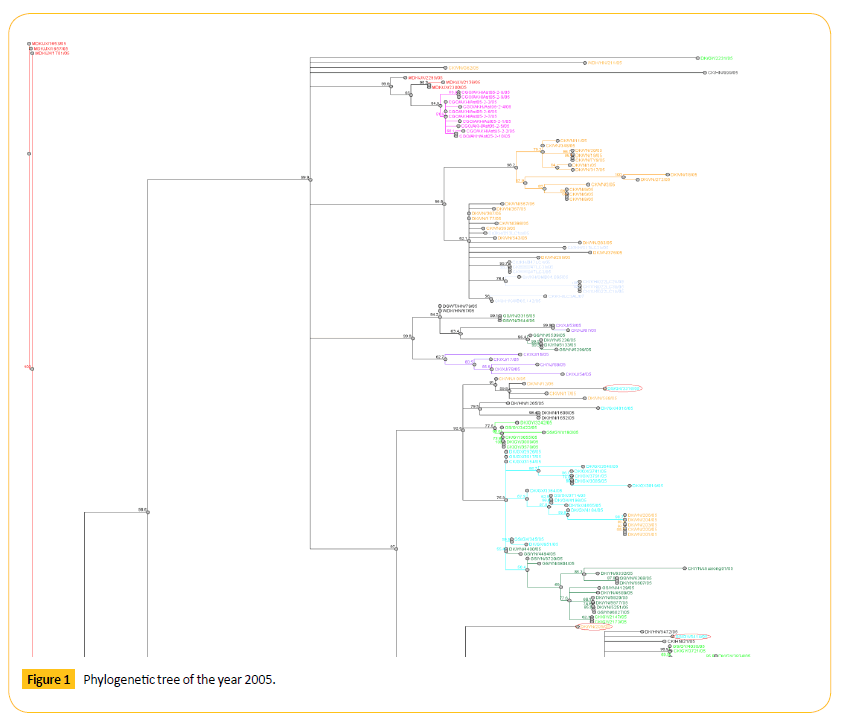
Figure 1: Phylogenetic tree of the year 2005.
In 2006 we have selected total 148 strains of Neuraminidase (NA) strain of H5N1 (Figure 2 and Table 2).
| Accession number |
Strain name |
| HQ200451 |
DK/KH/D14AL/06 |
| AB601142 |
CK/EGP/RIMD6-1/06 |
| EU889100 |
HGL/SE/V1116/06 |
| HM172177 |
DK/HN/29/06 |
| EF124273 |
GS/GX/532/06 |
Table 2: List of diverse strains from our analysis of 2006 are given below.
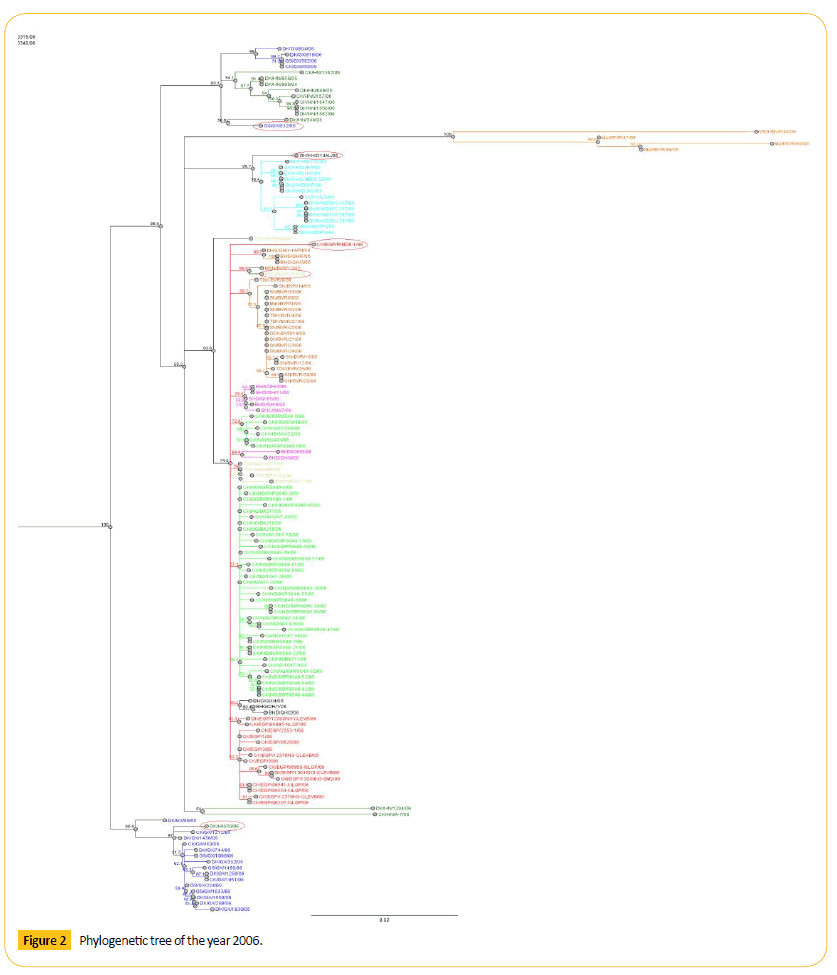
Figure 2: Phylogenetic tree of the year 2006.
In 2007 we have selected total 158 strains of Neuraminidase (NA) strain of H5N1 (Figure 3 and Table 3).
| Accession number |
Strain name |
| FJ602852 |
BHGS/QH/13/07 |
| FJ461727 |
BHGS/QH/F/07 |
| CY030455 |
DK/VN/NCVD-31/07 |
Table 3: List of diverse strains from our analysis of 2007 are given below.
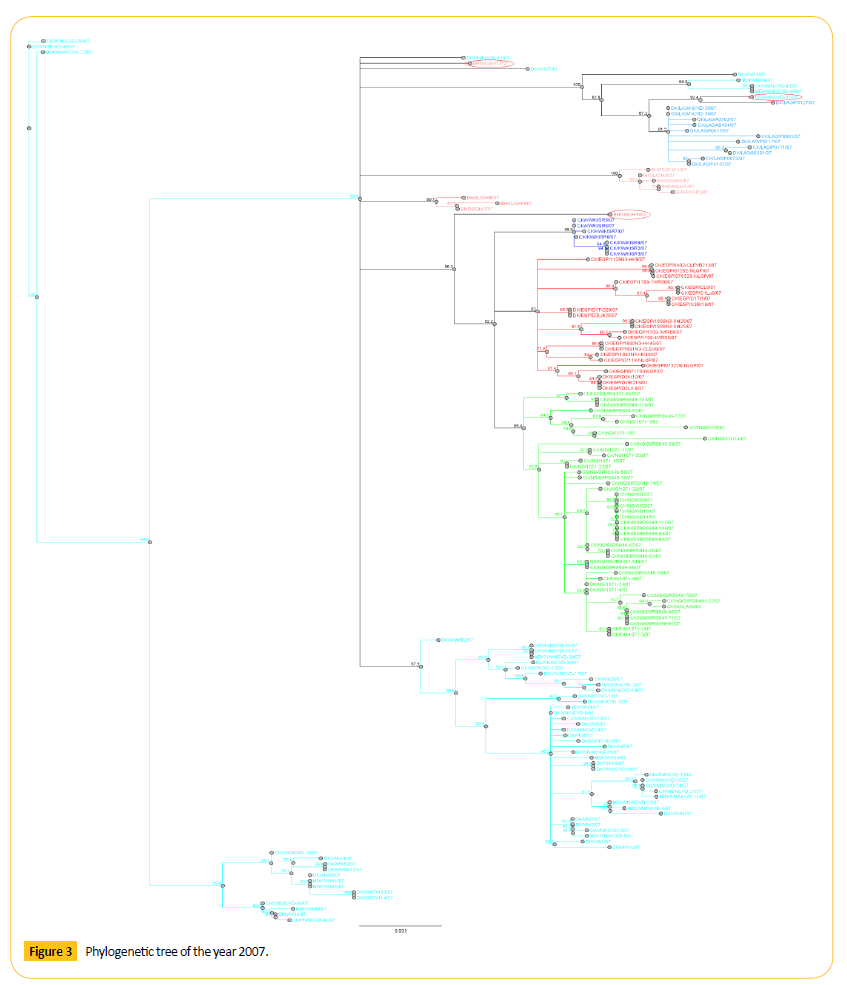
Figure 3: Phylogenetic tree of the year 2007.
From three years we have got some strains which seems to diverse from our analysis. We did our literature search but we did not get any information about these diverse strains. So it seems to us that these strains are not responsible for antigenic shift. Neighbor joining method and bootstrap value shows that this diverse strain is showing antigenic drift which may transfer to the other avian in the same country or other different country as well through migratory process.
From the phylogenetic tree of 2005 we came to see that all VN like viruses formed a cluster. Among them GS/GX/3316/05 shows 59.8 bootstrap value with DK/VN/12/05. Number of bootstrap value is showing the similarities between these two strains. Strain GS/GX/3316/05 may be migrated or may be mutated which can show antigenic drift.GS/GX/5414/05 shows 65.7 bootstrap values with DK/HN/5472/05. Number of bootstrap value is showing the similarities between these two strains. Strain GS/GX/5414/05 may be migrated or may be mutated which can show antigenic drift.
DK/JX/80/05 and WDK/HN/021/05 viruses are closely related to each other as they both are detached from their region and showing the same bootstrap value 69.8. One VN virus (DK/ VN/205/05) forming a clade with HB like viruses. Genetic character of this virus is different from other VN like viruses as this is detached from the cluster of VN like viruses showing the antigenic drift.
Phylogenetic tree analysis of 2006 shows that there are few diverse strains which may show genetic drift. One virus from Egypt (CK/EGP/RIMD6-1/06) form clade with the QG like viruses. HGL/ SE/V1116/06 shows 94.6 bootstrap values with MSN/BVR/12/06. Number of bootstrap value is showing the similarities between these two strains. Strain HGL/SE/V1116/06 may be migrated or may be mutated which can show antigenic drift. Among them GS/GX/532/06 shows 50.8 bootstrap values with DK/HN/344/06. Number of bootstrap value is showing the similarities between these two strains. Strain GS/GX/532/06 may be migrated or may be mutated which can show antigenic drift. One HN like virus (DK/ HN/29/06) forming a clade with Guangxi (GX) like viruses. One KH virus (DK/KH/D14AL/06) forming a clade. Genetic character of this virus is different from other KH like viruses as this is detached from the cluster of KH like viruses showing the antigenic drift.
Phylogenetic tree analysis of 2007 shows that there are few diverse strains which may show genetic drift. Among them DK/ VN/NCVD-31/07 shows 92.4 bootstrap values with DK/LAO/ PO127/07. Number of bootstrap value is showing the similarities between these two strains. Strain DK/VN/NCVD-31/07 may be migrated or may be mutated which can show antigenic drift. TWO QH virus (BHGS/QH/13/07 and BHGS/QH/F/07) forming a separate clade. Genetic character of this virus is different from other QH like viruses as these are detached from the cluster of QH like viruses showing the antigenic drift.
The molecular mechanisms that enable avian influenza viruses to cross the species barrier and transmit efficiently in humans are incompletely understood. Some experiments have been done to identify the transmission pattern and it shows poorer transmission from infected to susceptible animals [20-22]. Migration process can influence transmission of viruses. Migratory birds can carry pathogens from country to country thereby playing a role distributing influenza viruses. Avian influenza A consists of two major glycoproteins which are Hemagglutinin (HA) and Neuraminidase (NA) [23]. HA glycoproteins are more prone to attach to the cell surface sialic acid receptors. There is a difference between host surface receptors on the target cell which is believed to be the possible restrictive factor of avian influenza.
Human infections are periodic. In some cases these viruses are accompanied by high mortality. As a result they are the major concern about the potential H5N1 as an endemic virus.
Although human infections are sporadic; they are accompanied by high mortality; raising major concerns about the potential of H5N1 as a pandemic virus [24]. Fortunately; H5N1 viruses have not yet naturally acquired the ability to stably transmit between humans [25,26]. One factor that limits transmission of avian viruses in humans is the receptor specificity of the hemagglutinin (HA) [27]. Avian viruses; like H5N1; preferentially bind to α 2; 3 sialosides (avian-type receptors); whereas human viruses prefer α 2; 6 sialosides (human-type receptors that are found in the human respiratory tract).
Before functioning as a virus it needs post translational cleavage by host proteases [28]. HA followed by NA are important antigenic determinant from which neutralizing antibodies are directed. There are several subtypes of HA and NA. 18 different HA subtypes (H1 to H18) and 11 different NA subtypes (N1 to N11) are found [29].
In humans; the SA α 2;6 Gal receptor is expressed mainly in the upper airway; while the SA α 2;3 Gal receptor is expressed in alveoli and the terminal bronchiole [30]. A virus with good affinity to both SA α 2;3 Gal and SA α 2;6 Gal receptors may be a very dangerous one; which could both infect efficiently via its binding to SAα2; 6Gal in the upper airway and cause severe infection in the lung via its binding to SA α 2; 3Gal.
Data used in this study are obtained inside using nucleotide BLAST search from publicly available database of National Centre for Biotechnology Information (NCBI). Multiple sequence alignments; editing; assembly of strains were performed in windows platform with the Geneious program version 7.1.3 (trial).
In this study we will analyze some avian neuraminidase (H5N1) of different years. Analysis includes building nucleotide sequence and translating them into amino acid sequence. Then we will study amino acid positions with respect to some reported mutation to see the genetic pattern. After analyzing we will try to find out whether there are any similarities between avian and human or not. There are some reported avian H5N1 strains that affect human which are A/Goose/Hong Kong/739.2/2002 [31]; A/duck/Egypt/D1Br12/2007 [32]; A/Duck/Singapore/3/97 [33]; A/ egret/Egypt/1162/2006 [34]. All of these strain show preferential binding to Sia α (2; 6) Gal receptor that can infect a human. Few specific positions of amino acids are responsible for this binding. The NA of VN1203 in a PR8 background (designated N158D/ N224K/Q226L or N158D/N224K/Q226L/T318I; respectively) preferentially bind to Siaa2; 6Gal and attach to human tracheal epithelia [35,36] (Tables 4 and 5).
| Amino acid mutation position with reference |
Short form |
Amino acid in Geneious in reported position |
| Asn-158-Asp (Maineset al.) [34] |
N158D |
T |
| Asn-224-Lys (Chen et al.) [35] |
N224K |
W |
| Gln-226-Leu (Maineset al.) [34] |
Q226L |
N |
| Thr-318-Ile (Chen et al.) [35] |
T318I |
E |
Table 4: Summary of some reported position which can be responsible for antigenic shift from avian to human. Reported neuraminidase amino acid position with mutation that bind α 2, 6 receptor can affect human.
|
Reported AA with mutation point
|
AA in Geneious in reported position |
Exact AA which matches with the reported data with respect to specific mutation point (before 20 positions)
|
Exact AA which matches with the reported data with respect to specific mutation point (after 20 positions)
|
| N158D |
T158S(1) Polar to Polar T158A(2) Polar to Nonpolar |
N142 N147S(1) Polar to Polar |
N176 |
| N224K |
W224 |
N205S(7) Polar to Polar N205D(1) Polar to Negative N214 |
N226S(1) Polar to Polar N227Y(1) Polar to Polar N227S(1) Polar to Polar N240 |
| Q226L |
N226S(1) Polar to Polar |
|
Q232P(1) Polar to Nonpolar |
| T318I |
E318K(4) Electrically charged Negative to Positive |
|
T239A(1) Polar to Nonpolar T239K(1) Polar to Positive |
Table 5: Correlation of Reported neuraminidase amino acid position with mutation with our experimental strain, and their mutation pattern (exact and around twenty positions): Original Amino Acid analysis.
In case of N158D (Asn-158-Asp) reported position; we found amino acid Threonine (Thr) in our software (Geneious). We found two mutations here; Threonine (Thr) to Serine (Ser) in one strain and from Threonine (Thr) to Alanine (Ala) in two strains. We found amino acid Asparagine (Asn) in two positions (142 and 147) which is located within twenty positions before 158. We found one mutation here from Asparagine (Asn) to Serine (Ser) in one strain. We have also found Asparagine (Asn) in one position (176) which is located within twenty positions after 158. We found no mutation here. We found T158S and T158A mutation which indicates that polarity is changed from polar to polar and from polar to non-polar as Threonine (Thr); Serine (Ser) is Polar and Alanine (Ala) is non-polar.
In case of N224K (Asn-224-Lys) reported position; we found amino acid Tryptophan (Trp) in our software (Geneious). We found amino acid Asparagine (Asn) in two positions (205 and 214) which is located within twenty positions before 224. We found two mutations here from Asparagine (Asn) to Serine (Ser) in seven strains and from Asparagine (Asn) to Aspartic acid (Asp) in one strain. We have also found Asparagine (Asn) in three positions (226; 227 and 240) which is located within twenty positions after 224. We found three mutations here from Asparagine (Asn) to Serine (Ser) in one strain; Asparagine (Asn) to Tyrosine (Try) in one strain and from Asparagine (Asn) to Serine (Ser) in one strain. We found N205S and N205D mutation which indicates that polarity is changed from polar to polar and from polar to negative as Threonine (Thr); Serine (Ser) is Polar and Aspartic acid (Asp) is negative. We also found N227Y mutation which indicates that polarity is changed from polar to polar as Threonine (Thr); Serine (Ser) is Polar and Tyrosine is polar.
In case of Q226L (Gln-226-Leu) reported position; we found amino acid Threonine (Thr) in our software (Geneious). We found one mutations here; Threonine (Thr) to Serine (Ser) in one strain. We found amino acid Glutamine (Gln) in one position (232) which is located within twenty positions after 226. In this position there is one mutation here fron Glutamine (Gln) to Proline (Pro) in one strain. We found N226S and Q232P mutation which indicates that polarity is changed from polar to polar and from polar to non-polar as Threonine (Thr); Serine (Ser) is Polar and Proline (Pro) is non-polar.
In case of T318I (Thr-318-Ile) reported position; we found amino acid Glutamic acid (Glu) in our software (Geneious). We found one mutation here form Glutamiic (Glu) to Lysine (Lys) in four strain. We found amino acid Threonine (Thr) in position (239) which is located within twenty positions after 318. We found two mutations here from Threonine (Thr) to Alanine (Ala) in one strain and from Threonine (Thr) to Lysine (Lys) in one strain. In T239K; T239A and E318K mutations which indicate that polarity is changed from polar to polar; polar to positive and negative to polar as Threonine (Thr) is Polar; Alanine (Ala) is non-polar; Lysine (Lys) is positive and Glutamic acid (Glu) is negative (Table 6).
|
Reported AA with mutation point
|
AA in Geneious in reported position |
Exact AA which matches with the reported data with respect to specific mutation point (before 20 positions)
|
Exact AA which matches with the reported data with respect to specific mutation point (after 20 positions)
|
| N158D |
T158S(1) Polar to Polar T158A(2) Polar to Nonpolar |
D143 D152 |
|
| N224K |
W224 |
K212 |
|
| Q226L |
N226S(1) Polar to Polar |
L211V(1) Nonpolar to Nonpolar |
L229 |
| T318I |
E318K(4) Electrically charged Negative to Positive |
|
I321T(1) Nonpolar to Polar I321M(3) Nonpolar to Nonpolar I321V(2) Nonpolar to Nonpolar I324 |
Table 6: Correlation of Reported neuraminidase amino acid position with mutation with our experimental strain, and their mutation pattern (exact and around twenty positions): Mutated Amino Acid analysis.
In case of N158D (Asn-158-Asp) reported position; we found amino acid Threonine (Thr) in our software (Geneious). We found two mutations here; Threonine (Thr) to Serine (Ser) in one strain and from Threonine (Thr) to Alanine (Ala) in two strains. We found amino acid Asparagine (Asn) in two positions (143 and 152) which is located within twenty positions before 158. There is no mutation here. We found T158S and T158A mutation which indicates that polarity is changed from polar to polar and from polar to non-polar as Threonine (Thr); Serine (Ser) is polar and Alanine (Ala) is non-polar.
In case of N224K (Asn-224-Lys) reported position; we found amino acid Tryptophan (Trp) in our software (Geneious). We found amino acid Lysine (Lys) in one positions (212) which is located within twenty positions before 224. There is no mutation here.
In case of Q226L (Gln-226-Leu) reported position; we found amino acid Threonine (Thr) in our software (Geneious). We found one mutations here; Threonine (Thr) to Serine (Ser) in one strain. We found amino acid Leucine (Leu) in one position (211) which is located within twenty positions before 226. . In this position there is one mutation here fronLeucine (Leu) to Valine (Val) in one strain. We found amino acid Leucine (Leu) in one position (229) which is located within twenty positions after 226. There is no mutation hereWe found N226S and L211V mutation which indicates that polarity is changed from polar to polar and from non-polar to non-polar as Threonine (Thr); Serine (Ser) is Polar and Proline (Pro); Valine (Val) is non-polar.
In case of T318I (Thr-318-Ile) reported position; we found amino acid Glutamic acid (Glu) in our software (Geneious). We found one mutation here form Glutamiic (Glu) to Lysine (Lys) in four strains.
We found amino acid Isoleucine (Ile) in two positions (321 and 324) which is located within twenty positions after 318. We found three mutations here from Isoleucine (Ile) to Threonine (Thr) in one strain; from Isoleucine (Ile) to Methionine (Met) in three strains and from Isoleucine (Ile) to Valine (Val) in two strains. In I321T; I321M; I321V and E318K mutations which indicate that polarity is changed from non-polar to non-polar and non-polar to polar as Isoleucine (Ile); Methionine (Met) and Valine (Val) is nonpolar and Threonine (Thr) is polar (Tables 7 and 8).
|
Amino acid mutation position with reference
|
Short form
|
Amino acid in Geneious in reported position
|
NAIs |
His-274-Try
(Le QMet al.) [36] |
H274Y |
V |
Oseltamivir Peramivir |
His-275-Try
(Pizzornoet al.) [38] |
H275Y |
E |
Oseltamivir Peramivir |
Asn-294-Ser
(Le QMet al.) [36] |
N294S |
E |
Oseltamivir |
Asn-295-Ser
(Pizzornoet al.) [38] |
N295S |
I |
Oseltamivir |
Asp-198-Gly
(Hurt et al.) [37] |
D198G |
I |
Oseltamivir Zanamivir |
Asp-199-Gly
(Pizzornoet al.) [38] |
D199G |
G |
Oseltamivir Zanamivir |
Val -116- Ala
(Nguyen et al.) [39] |
V116A |
V |
Oseltamivir Zanamivir |
His -252- Try
(Nguyen et al.) [39] |
H252Y |
S |
Oseltamivir |
Ser -246- Asn
(Nguyen et al.) [39] |
S246N |
V |
Oseltamivir |
Ile -222- Thr
(Nguyen et al.) [39] |
I222T |
K |
Oseltamivir |
Table 7: Summary of some reported position which can be responsible for N1 (H5N1) neuraminidase mutations causing reduced sensitivity to NAIs [37-39].
| Reported AA with mutation point |
AA in Geneious in reported position |
Exact AA which matches with the reported data with respect to specific mutation point
(before 20 positions) |
Exact AA which matches with the reported data with respect to specific mutation point
(after 20 positions) |
| H274Y |
V274I (2)
Nonpolar to Nonpolar |
|
H282 |
| H275Y |
E275 |
|
H282 |
| N294S |
E294G
Electrically charged
Negative to Nonpolar |
N280D(1)
Polar to Electrically charged Negative
N280Y(1)
Polar to Polar
N280T(1)
Polar to Polar
N280K(1)
Polar to Electrically
Charged Positive |
N302
N307
N314 |
| N295S |
I295V(7)
Nonpolar to Nonpolar |
N280D(1)
Nonpolar to Electrically charged
Negative
N280Y(1)
Polar to Polar
N280T(1)
Polar to Polar
N280K(1)
Polar to Electrically
Charged Positive |
N302
N307
N314 |
| D198G |
I198 |
D191 |
D204 |
| D199G |
G199 |
D191 |
D204
D219 |
| V116A |
V116I(2)
Nonpolar to Nonpolar |
V99 I
Nonpolar to Nonpolar
V114 |
|
| H252Y |
S252N(4)
Polar to Polar
S 252 G(4)
Polar to Nonpolar |
|
|
| S246N |
V246L(3)
V246I(6)
Nonpolar to Nonpolar (L,I) |
S 234
S 242 F(1)
Polar to Nonpolar |
S252G(4)
Polar to Nonpolar
S252N(4)
Polar to Polar
S257 |
| I222T |
K222Q(1)
K222R(3)
K222N(1)
Electrically charged
Positive to Polar
(Q, N)and to Positive (R) |
I216V(1)
I217
I221V(1)
Nonpolar to Nonpolar |
I228V(1)
I228T(2)
I228L(1)
Nonpolar to Nonpolar (V,L) and to Polar (T) |
Table 8: Analysis of our study data with respect to reported mutation point causing reduced sensitivity to NAIs of avian H5N1. Correlation of Reported NAIs resistance specific avian amino acid position with our experimental strain, and their mutation pattern (exact and around twenty positions): Original Amino Acid analysis.
In case of H274Y (His-274-Try) reported position; we found amino acid Valine (V) in our software (Geneious). We found one mutation here; Valine (V) to Isoleucine (I) in two strains. We found amino acid Histidine (H) in one positions (282) which are located after twenty positions 274. We found V 274 I (2) mutation in two strains which indicates that there is polarity changed from non-polar to non-polar as Valine (V) is non-polar and Isoleucine (I) is non-polar.
In case of H275Y (His-274-Try) reported position; we found amino acid Glutamic acid (E) in our software (Geneious). There are no mutations here. We found amino acid Histidine (H) in one positions (282) which are located after twenty positions 274. There are no mutations here.
In case of N294S (Asn-294-Ser) reported position; we found amino acid Glutamic acid (E) in our software (Geneious). We found one mutation here; Glutamic acid (E) to Glycine (G) in one strain. We found Asparagine (Asn) in one position (280) which is located within twenty positions before 294. In 280 positions there are four mutations here from Asparagine (N) to Aspartic acid (D) in one strain; from Asparagine (N) to Tyrosine (Y) in one strain; from Asparagine (N) to Threonine (T) in one strain and from Asparagine (N) to Lysine (K) in one strain. We found amino acid Asparagine (N) in three positions (302; 307 and 314) which are located after twenty positions 294. There are no mutation here. In N280D; N280Y; N280T and N280K mutations which are indicate that there is polarity changed from polar to electrically charged negative; to positive and to polar as Asparagine (N) is polar; Aspartic acid (D) is electrically charged negative; Tyrosine (Y) is polar; Threonine (T) is polar and Lysine (K) is electrically charged positive. In E294G mutation which is indicates that there is polarity change from electrically charged negative to nonpolar as Glutamic acid (E) is negative and Glycine (G) is nonpolar.
In case of N295S (Asn-295-Ser) reported position; we found amino acid Isoleucine (I) in our software (Geneious). We found one mutation here; Isoleucine (I) to Valine (V) in seven strains.
We found Asparagine (Asn) in one position (280) which is located within twenty positions before 295. In 280 positions there are four mutations here from Asparagine (N) to Aspartic acid (D) in one strain; from Asparagine (N) to Tyrosine (Y) in one strain; from Asparagine (N) to Threonine (T) in one strain and from Asparagine (N) to Lysine (K) in one strain. We found amino acid Asparagine (N) in three positions (302; 307 and 314) which are located after twenty positions 295.There are no mutation here. We found I295V mutation which indicates that there is polarity changed from non-polar to non-polar as Isoleucine (I) is non-polar and Valine (V) is non-polar. In N280D; N280Y; N280T and N280K mutations which are indicate that there is polarity changed from polar to electrically charged negative; to positive and to polar as Asparagine (N) is polar; Aspartic acid (D) is electrically charged negative; Tyrosine (Y) is polar; Threonine (T) is polar and Lysine (K) is electrically charged positive.
In case of D198G (Asp-198-Gly) reported position; we found amino acid Isoleucine (I) in our software (Geneious). There are no mutations here. We found amino acid Aspartic acid (D) in one positions (191) which are located before twenty positions 198. We found amino acid Aspartic acid (D) in one positions (204) which are located after twenty positions 198. There are no mutations here.
In case of D199G (Asp-199-Gly) reported position; we found amino acid Glycine (G) in our software (Geneious). There are no mutations here. We found amino acid Glycine (G) in one positions (191) which is located within twenty positions before 199. There are no mutations here. We found amino acid Aspartic acid (D) in two positions (204 and 219) which are located after twenty positions 199. There are no mutations here.
In case of V116A (Val-116-Ala) reported position; we found amino acid Valine (V) in our software (Geneious). There is one mutation here from Valine (V) to Isoleucine (I) in two strains. We found amino acid Valine (V) in two positions (99 and 114) which is located within twenty positions before 116 and one mutation here from Valine (V) to Isoleucine (I) in one strain. We found V116I mutation which is indicates that there is polarity changed from non-polar to non-polar as Valine (V) is non-polar and Isoleucine (I) is non-polar.
In case of H252Y (His-252-Try) reported position; we found amino acid Serine (S) in our software (Geneious). There are two mutations here from Serine (S) to Asparagine (N) in four strains and from Serine (S) to Glycine (G) in four strains. We found S252N mutation which is indicates that there is polarity changed from polar to polar as Serine (S) is polar and Asparagine (N) is polar. We found S252G mutation which is indicates that there is polarity changed from Polar to non-polar as Serine (S) is polar and Glycine (G) is non-polar.
In case of S246N (Ser-246-Asn) reported position; we found Valine (V) in our software (Geneious). There are two mutations here from Valine (V) to Leucine (L) in three strains and from Valine (V) to Isoleucine (I) in six strains. We found V246L and V246I mutation which is indicates that there is polarity changed from non-polar to non-polar as Valine (V) is non-polar; Leucine (L) is non-polar and Isoleucine (I) is non-polar. We found amino acid Serine (S) in two positions (234 and 242) which is located within twenty positions before 246. In 234 positions there is no mutation here. In 242 positions there is one mutation from Serine (S) to Phenylalanine (F) in one strain.
We found S242F mutation which is indicates that there is polarity changed from polar to non-polar as Serine (S) is polar and Phenylalanine (F) is non-polar. We found amino acid Serine (S) in two positions (252 and 257) which is located within twenty positions after 246. In 252 positions there are two mutations here from Serine (S) to Glycine (G) in four strains and from Serine (S) to Asparagine (N) in four strains. In 257 positions there is no mutation here. We found S252G mutation which is indicates that there is polarity changed from polar to non-polar as Serine (S) is polar and Glycine (G) is non-polar. We found S252N mutation which is indicates that there is polarity changed from Polar to Polar as Serine (S) is polar and Asparagine (N) is polar.
In case of I222T (Ile-222-Thr) reported position; we found Lysine (K) in our software (Geneious). There are three mutations here from Lysine (K) to Glutamine (Q) in one strain; from Lysine (K) to Arginine (R) in three strains and from Lysine (K) to Asparagine (N) in one strain. We found K222Q; K222R and K222N mutation which is indicates that there is changed electrically charged from positive (K) to positive (R) and from positive (K) to polar (Q and N).
We found amino acid Isoleucine (I) in three positions (216; 217 and 221) which is located within twenty positions before 222. In 216 positions there is one mutation here from Isoleucine (I) to Valine (V) in one strain and in 221 positions there is one mutation here from Isoleucine (I) to Valine (V) in one strain. In 217 positions there is no mutation here. In I216V and I221V mutations which are indicate that there is polarity changed from non-polar to nonpolar as Isoleucine (I) is non-polar and Valine (V) is non-polar.
We found amino acid Isoleucine (I) in one positions (228) which are located within twenty positions after 222. In 228 positions there are three mutations here from Isoleucine (I) to Valine (V) in one strain; from Isoleucine (I) to Threonine (T) in two strains and from Isoleucine (I) to Leucine (L) in one strain. In I228V; I228T and I228L mutations which are indicate that there is polarity changed from non-polar to non-polar and non-polar to polar as Isoleucine (I) is non-polar; Valine (V) is non-polar; Leucine (L) is non-polar and Threonine (T) is polar (Table 9).
|
Reported AA with mutation point
|
AA in Geneious in reported position
|
Exact AA which matches with the reported data with respect to specific mutation point
(before 20 positions)
|
Exact AA which matches with the reported data with respect to specific mutation point
(after 20 positions)
|
| H274Y |
V274I(2)
Nonpolar to Nonpolar |
Y258 |
Y281F(2)
Polar toNonpolar
Y283
Y289N(1)
Polar to Polar |
| H275Y |
E275 |
Y258 |
Y281F(2)
Polar toNonpolar
Y283
Y289N(1)
Polar to Polar |
| N294S |
E294G
Electrically charged
Negative to Nonpolar |
S287 |
S306
S312 |
| N295S |
I295V(7)
Nonpolar to Nonpolar |
S287 |
S306
S312 |
| D198G |
I198 |
G192 |
G199
G202
G206A(1)
Nonpolar to Nonpolar
G206E(2)
Nonpolar to Electrically
Charged Negative
G215 |
| D199G |
G199 |
G 192 |
G202
G206A(1)
Nonpolar to Nonpolar
G206E(2)
Nonpolar to Electrically
Charged Negative
G215 |
| V116A |
V11I(2)
Nonpolar to Nonpolar |
A98 |
|
| H252Y |
S252N(4)
Polar to Polar
S252G(4)
Polar to Nonpolar |
|
Y258 |
| S246N |
V246L(3)
V246I(6)
Nonpolar to Nonpolar (L,I) |
N240 |
N253D(1)
Polar to Electrically charged Negative
N253H(1)
Polar to Electrically charged Positive
N253S(2)
Polar to Polar |
| I222T |
K222Q(1)
K222R(3)
K222N(1)
Electrically charged
Positive to Polar
(Q,N)and to Positive (R) |
T220
T218 |
T231 |
Table 9: Correlation of Reported NAIs resistance specific avian amino acid position with our experimental strain, and their mutation pattern (exact and around twenty positions): Mutated amino acid analysis.
In case of H274Y (His-274-Try) reported position; we found amino acid Valine (V) in our software (Geneious). We found one mutation here; Valine (V) to Isoleucine (I) in two strains. We found amino acid Tyrosine (Y) in one position (258) which is located before twenty positions 274. There are no mutations here. We found amino acid Tyrosine (Y) in three positions (281; 283 and 289) which are located after twenty positions after 274. There are two mutations here from Tyrosine (Y) to Phenylalanine (F) in two strains and from Tyrosine (Y) to Asparagine (N) in one strain. We found V274I mutation in two strains which indicates that there is polarity changed from non-polar to non-polar as Valine (V) is non-polar and Isoleucine (I) is non-polar. We found Y281F and Y289N mutations which indicate that there is polarity changed from polar to non-polar and to polar as Tyrosine (Y) is polar; Phenylalanine (F) is non-polar and Asparagine (N) is polar.
In case of H275Y (His-274-Try) reported position; we found amino acid Glutamic acid (E) in our software (Geneious). We found amino acid Tyrosine (Y) in one position (258) which is located before twenty positions 275. There are no mutations here. We found amino acid Tyrosine (Y) in three positions (281; 283 and 289) which are located after twenty positions after 275. There are two mutations here from Tyrosine (Y) to Phenylalanine (F) in two strains and from Tyrosine (Y) to Asparagine (N) in one strain. We found Y281F and Y289N mutations which indicate that there is polarity changed from polar to non-polar and to polar as Tyrosine (Y) is polar; Phenylalanine (F) is non-polar and Asparagine (N) is polar.
In case of N294S (Asn-294-Ser) reported position; we found amino acid Glutamic acid (E) in our software (Geneious). We found one mutation here; Glutamic acid (E) to Glycine (G) in one strain. We found amino acid Serine (S) in one position (287) which is located within twenty positions before 294. We found amino acid Serine (S) in two positions (306 and 312) which is located within twenty positions after 294. There are no mutations here. In E294G mutation which is indicates that there is polarity change from electrically charged negative to nonpolar as Glutamic acid (E) is negative and Glycine (G) is nonpolar.
In case of N295S (Asn-295-Ser) reported position; we found amino acid Isoleucine (I) in our software (Geneious). We found one mutation here; Isoleucine (I) to Valine (V) in seven strains. We found amino acid Serine (S) in one position (287) which is located within twenty positions before 295. We found amino acid Serine (S) in two positions (306 and 312) which is located within twenty positions after 295. There are no mutations here. We found I295V mutation which indicates that there is polarity changed from non-polar to non-polar as Isoleucine (I) is non-polar and Valine (V) is non-polar.
In case of D198G (Asp-198-Gly) reported position; we found amino acid Isoleucine (I) in our software (Geneious). There are no mutations here. We found amino acid Glycine (G) in one position (192) which is located within twenty positions before 198. We found amino acid Glycine (G) in four positions (199; 202; 206 and 215) which is located within twenty positions after 198. We found two mutations here from Glycine (G) to Alanine (A) in one strain and from Glycine (G) to Glutamic acid (E) in two strains. In G206A and G206E mutations which indicates that there is polarity changed from nonpolar to nonpolar and nonpolar to electrically charge negative as Glycine (G) and Alanine (A) is nonpolar and Glutamic acid (E) is electrically charged negative.
In case of D199G (Asp-198-Gly) reported position; we found amino acid Isoleucine (I) in our software (Geneious). There are no mutations here. We found amino acid Glycine (G) in one position (192) which is located within twenty positions before 199. We found amino acid Glycine (G) in three positions (202; 206 and 215) which is located within twenty positions after 199. We found two mutations here from Glycine (G) to Alanine (A) in one strain and from Glycine (G) to Glutamic acid (E) in two strains. In G206A and G206E mutations which indicates that there is polarity changed from nonpolar to nonpolar and nonpolar to electrically charge negative as Glycine (G) and Alanine (A) is nonpolar and Glutamic acid (E) is electrically charged negative.
In case of V116A (Val-116-Ala) reported position; we found amino acid Valine (V) in our software (Geneious). There is one mutation here from Valine (V) to Isoleucine (I) in two strains. We found amino acid Alanine (A) in one position (98) which is located within twenty positions before 116. There are no mutations here. We found V116I mutation which is indicates that there is polarity changed from non-polar to non-polar as Valine (V) is non-polar and Isoleucine (I) is non-polar.
In case of H252Y (His-252-Try) reported position; we found amino acid Serine (S) in our software (Geneious). There are two mutations here from Serine (S) to Asparagine (N) in four strains and from Serine (S) to Glycine (G) in four strains. We found S252N mutation which is indicates that there is polarity changed from polar to polar as Serine (S) is polar and Asparagine (N) is polar. We found amino acid Tyrosine (Y) in one position (258) which is located within twenty positions after 252. There are no mutations here. We found S252G mutation which is indicates that there is polarity changed from Polar to non-polar as Serine (S) is polar and Glycine (G) is non-polar.
In case of S246N (Ser-246-Asn) reported position; we found Valine (V) in our software (Geneious). There are two mutations here from Valine (V) to Leucine (L) in three strains and from Valine (V) to Isoleucine (I) in six strains. We found amino acid Asparagine (N) in one position (240) which is located within twenty positions before 246. There is no mutation here. We found amino acid Asparagine (N) in one position (253) which is located within twenty positions after 246. There are three mutations here from Asparagine (N) to Aspartic acid (D) in one strain; from Asparagine (N) to Histidine (H) in one strain and fron Asparagine (N) to Serine (S) in two strains. We found N253D; N253H and N253S mutation which is indicates that there is polarity changed from polar to electrically charged negative; to positive and to polar as Asparagine (N) is polar; Aspartic acid (D) is electrically charged negative; Histidine (H) is electrically charged positive and Serine (S) is polar. We found V246L and V246I mutation which is indicates that there is polarity changed from non-polar to non-polar as Valine (V) is non-polar; Leucine (L) is non-polar and Isoleucine (I) is non-polar.
In case of I222T (Ile-222-Thr) reported position; we found Lysine (K) in our software (Geneious). There are three mutations here from Lysine (K) to Glutamine (Q) in one strain; from Lysine (K) to Arginine (R) in three strains and from Lysine (K) to Asparagine (N) in one strain. We found amino acid Threonine (T) in two positions (220 and 218) which is located within twenty positions before 222. We found amino acid Threonine (T) in one positions (231) which is located within twenty positions after 222.There are no mutations here. We found K222Q; K222R and K222N mutation which is indicates that there is changed electrically charged from positive (K) to positive (R) and from positive (K) to polar (Q and N).
Conclusion
Neuraminidase inhibitors (NAIs) are first-line agents for the treatment and prevention of influenza virus infections. Resistance to the NAIs can be both drug and virus type or subtype specific. The effects of NAI resistance NA mutations on the fitness and transmissibility of influenza viruses may vary depending on several factors: location of the mutation (catalytic or framework residue); NA type ⁄ subtype; virus genetic background; existence of permissive secondary NA mutations; degree of NA functional loss; and an appropriate functional NA–HA balance. In addition; differences in the host’s immune response and genetic background can contribute to such variation. No single measure can easily describe the extent of fitness of influenza viruses carrying drug resistance mutations.
The most frequently reported change conferring oseltamivir resistance in that viral context is the H274Y neuraminidase mutation (N1 numbering). Recent studies have shown that; in the presence of the appropriate permissive mutations; the H274Y variant can retain virulence and transmissibility in some viral backgrounds. Most oseltamivir-resistant influenza A virus infections can be managed with the use of inhaled or intravenous zanamivir; another NAI. New NAI compounds and non-neuraminidase agents as well as combination therapies are currently in clinical evaluation for the treatment for severe influenza infections.
At present; it is clear that mutation conferring resistance to the currently approved antiviral drugs is a growing problem; so it is very important to continue research in various areas that would allow better resolution of the problem: First; the knowledge of mutation’s type that allows virus avoids the action of drugs; will allow an understanding as to how these mutations arise and how we must avoid them. Also; this knowledge; will allow a better design of new drugs.
Conflict of Interest
None
8005
References
- Matrosovich MN, Matrosovich TY, Gray T, Roberts NA, Klenk HD (2004) Neuraminidase is important for the initiation of influenza virus infection in human airway epithelium. Journal of Virology 78: 12665-12667.
- Suzuki T, Takahashi T, Guo CT, Hidari KIPJ, Miyamoto D, et al. (2005)Sialidase activity of influenza A virus in an endocytic pathway enhances viral replication. Journal of Virology 79: 11705-11715.
- Yen HL, Hoffmann E, Taylor G, Scholtissek C, Monto AS, et al. (2006) Importance of neuraminidase active-site residues to the neuraminidase inhibitor resistance of influenza viruses. Journal of Virology 80: 8787-8795.
- Garcia J, Sovero M, Torres AL, Gomez J, Douce R, et al. (2009) Antiviral resistance in influenza viruses circulating in Central and South America based on the detection of established genetic markers. Influenza and Other Respiratory Viruses 3: 69-74.
- Cao B, Wang DY, Yu XM, We LQ, Pu ZH, et al. (2012) An uncontrolled openlabel, multicenter study to monitor the antiviral activity and safety of inhaled zanamivir (as Rotadisk via Diskhaler device) among Chinese adolescents and adults with influenza-like illness. Chinese Medical Journal 125: 3002-3007.
- Ito T, Goto H, Yamamoto E, Tanaka H, Takeuchi M, et al. (2001) Generation of a highly pathogenic avian influenza A virus from an avirulent field isolate by passaging in chickens. J Virol 75:4439-4443.
- Matrosovich M, Zhou N, Kawaoka Y, Webster R (1999) The Surface Glycoproteins of H5 Influenza Viruses Isolated from Humans, Chickens, and Wild Aquatic Birds Have Distinguishable Properties. Journal of Virology 73: 1146-1155.
- Rott R (1992) The pathogenic determinant of influenza virus. Vet Microbiol 33: 303-310.
- Tong S, Zhu X, Li Y, Shi M, Zhang J, et al. (2013) New world bats harbor diverse influenza A viruses. PLoSPathog 9: e1003657.
- Jing X, MaC,Ohigashi Y, Oliveira FA, Jardetzky TS, et al. (2008) Functional studies indicate amantadine binds to the pore of the influenza A virus M2 proton-selective ion channel. Proc. Natl. Acad. Sci. U.S.A105: 10967-10972.
- Mendel DB, Roberts NA (1998) In-vitro and in-vivo efficacy of influenza neuraminidase inhibitors. CurrOpin Infect Dis 11: 727-732.
- DgxncghSK, Mitamura K, Sakai TY, Goto H, Sugaya N, et al. (2003) High frequency of resistant viruses harboring different mutations in amantadine-treated children with influenza. J Infect Dis 188:57-61.
- Russell RJ, Haire LF, Stevens DJ, Collins PJ, Lin YP, et al. (2006) The structure of H5N1 avian influenza neuraminidase suggests new opportunities for drug design. Nature 443: 45-49.
- Van der VE, Schutten M, Boucher CA (2011) The potential for multidrug-resistant influenza. Current Opinion Infectious Diseases 24: 599-604.
- De Jong M, Thanh TT, Khanh TH (2005) Oseltamivir resistance during treatment of influenza A (H5N1) infection. N Engl J Med 353:2667-2672.
- Le QM, Kiso M, Someya K, Sakai YT, Nguyen TH, et al. (2005) Avian flu: isolation of drug-resistant H5N1 virus. Nature 437: 1108.
- Earhart KC, Elsayed NM, Saad MD (2009) Oseltamivir resistance mutation N294S in human influenza A(H5N1) virus in Egypt. J Infect Public Health 2:74-80.
- Le QM, Kiso M, Someya K, Sakai YT, Nguyen TH, et al. (2005) Avian flu: isolation of drug-resistant H5N1 virus. Nature 437: 1108.
- Narayan O, Lang G, Rouse ST (1969) A new influenza A virus infection in turkeys. IV Experimental susceptibility of domestic birds to virus strain ty/Ontario/7732/1966.Archivfurdiegesamte Virus forschung26:149-165.
- Alexander DJ, Allan WH, Parsons D, Parsons G (1978) The pathogenicity of four avian influenza viruses for fowls, turkeys and ducks.Research in Veterinary Science24:242-247.
- Westbury HA, Turner AJ, Kovesdy L (1979)The pathogenicity of three Australian fowl plague viruses for chickens, turkeys and ducks. Veterinary Microbiology 4:223-234.
- Ito T, Goto H, Yamamoto E, Tanaka H, Takeuchi M, et al. (2001) Generation of a highly pathogenic avian influenza A virus from an avirulent field isolate by passaging in chickens. J Virol 75: 4439-4443.
- Gambotto A, Barratt-Boyes SM, de Jong MD, Neumann G, Kawaoka Y (2008) Human infection with highly pathogenic H5N1 influenza virus. Lancet371:1464-1475.
- Ungchusak K, Auewarakul P, Dowell SF, Kitphati R, Auwanit W, et al. (2005) Probable person-to-person transmission of avian influenza A (H5N1). N. Engl. J. Med352:333-340.
- Wang H, Feng Z, Shu Y, Yu H, Zhou L, et al. (2008) Probable limited person-to-person transmission of highly pathogenic avian influenza A (H5N1) virus in China. Lancet 371:1427-1434.
- Imai M, Kawaoka Y (2012) The role of receptor binding specificity in interspecies transmission of influenza viruses. CurrOpinVirol 2: 160-167.
- Rott R (1992) The pathogenic determinant of influenza virus. Vet Microbiol 33: 303-310.
- Tong S, Zhu X, Li Y, Shi M, Zhang J, et al. (2013) New World Bats Harbor Diverse Influenza A Viruses. PLoS Pathogens9: e1003657.
- Shinya K, Ebina M, Yamada S, Ono M, Kasai N, et al. (2006) Avian flu: influenza virus receptors in the human airway. Nature 440: 435-436.
- Gambaryan A, Tuzikov A, Pazynina G, Bovin N, Balish A, et al. (2006) Evolution of the receptor binding phenotype of influenza A (H5) viruses. Virology344:432-438.
- Yohei W, Madiha SI, Hany FE, Norihito K, Rika M, et al. (2011) Acquisition of Human-Type Receptor Binding Specificity by New H5N1 Influenza Virus Sublineages during Their Emergence in Birds in Egypt. PLoS Pathogens7: e1002068.
- Nipa J, Wanwimon M, Daungmanee C, Philip JS, Sissades T, et al. (2009) Prediction of avian influenza A binding preference to human receptor using conformational analysis of receptor bound to hemagglutinin. BMC Genomics10:S24.
- De Vries RP, Zhu X, McBride R, Rigter A, Hanson A, et al. (2014) Hemagglutinin Receptor Specificity and Structural Analyses of Respiratory Droplet-Transmissible H5N1 Viruses. Journal of Virology88:768-773.
- Maines TR, Chen LM, Van Hoeven N, Tumpey TM, Blixt O, et al. (2011) Effect of receptor binding domain mutations on receptor binding and transmissibility of avian influenza H5N1 viruses. Virology 413: 139-147.
- Chen LM (2012) In vitro evolution of H5N1 avian influenza virus toward humantype-receptor specificity. Virology 422: 105-113.
- Le QM, Kiso M, Someya K, Sakai YT, Nguyen TH, et al. (2005) Avian flu: isolation of drug-resistant H5N1 virus. Nature 437: 1108.
- Hurt AC, Holien JK, Parker M, Kelso A, Barr IG (2009) Zanamivir-resistant influenza viruses with a novel neuraminidase mutation. J Virol 83: 10366-10373.
- Pizzorno A, Bouhy X, Abed Y, Boivin G (2011) Generation and characterization of recombinant pandemic influenza A(H1N1) viruses resistant to neuraminidase inhibitors. J Infect Dis 203: 25-31.
- Nguyen HT, Fry AM, Loveless PA, Klimov AI, Gubareva LV (2010) Recovery of a multidrugresistant strain of pandemic influenza A 2009 (H1N1) virus carrying a dual H275Y/I223R mutation from a child after prolonged treatment with oseltamivir. Clin Infect Dis 51: 983-984.