Magellan Guewo Fokeng1,2*, Barbara Atogho-Tiedeu1,2, Eugene Sobngwi2,3, Jean-Claude Mbanya2,3 and Wilfred Fon Mbacham2,4
1Department of Biochemistry, Faculty of Science, University of Yaounde I, Yaounde, Cameroon
2The Biotechnology Centre, University of Yaoundé I, Yaoundé, Cameroon
3Department of Internal Medicine, Faculty of Medicine and Biomedical Sciences, University of Yaounde I, Yaounde, Cameroon
4Department of Biochemistry, Physiology and Pharmacology, Faculty of Medicine and Biomedical Sciences, University of Yaounde I, Yaounde, Cameroon
- Corresponding Author:
- Prof. Wilfred Fon Mbacham
Department of Biochemistry, Physiological Science and Pharmacology, Faculty of Medicine and Biomedical Sciences, University of Yaoundé I, Yaoundé, Cameroon
Tel: +236 75767086
E-mail: wfmbacham@yahoo.com
Received date: Apr 29, 2016; Accepted date: May 26, 2016; Published date: May 30, 2016
Citation: Fokeng MG, Atogho-Tiedeu B, Sobngwi E, et al. The Krüppel-Like Factor 14 (KLF14), Master gene of multiple metabolic phenotypes: Putative Trans-Regulator Network. Transl Biomed. 2016, 7:2.
Keywords
Type 2 diabetes; Kruppel-like factor 14 (KLF14); Transcription factor; Gene regulation
Background
The prevalence of type 2 diabetes mellitus (T2DM) has increased rapidly not only in affluent societies, but also in developing countries over the last 20 years [1]. This indicates that there is a global health crisis stemming from changing life styles. Worldwide, there are more than 415 million with diabetes which are projected to rise to 642 million by 2040 [2]. The increasing global prevalence of T2DM is also tied to rising rates of obesity [3]. It is commonly said that diabetes runs in the family because people do not run and points to these diseases as being multifactorial in which environmental triggers interact with genetic variants in the predisposition to the disease. The discovery of causal genes has followed three main waves. The first wave consisted of family-based linkage analyses and focused candidate-gene studies [4], the second wave of discovery involved a switch to tests of association [5], and the third, and most successful wave of discovery has been driven by systematic, large-scale surveys of association between common DNA sequence variants and disease [6]. McCarthy showed in a review on “Genomics, Type 2 Diabetes, and Obesity” that there were 67 (sixty seven) genomic locations of proven signals of non-autoimmune forms of diabetes and 62 (sixty two) genomic locations of proven signals of body-mass index, obesity, and related phenotypes [7].
In 2011, Kerrin and collaborators focused on ten genes (TPMT, ARSD, SLC7A10, C8 or f82, APH1B, PRMT2, NINJ2, KLF13, GNB1, and MYL5) showing genome-wide significant trans (GWST) associations that was driven by rs4731702, leading to the identification of KLF14 gene as a master transregulator related to multiple metabolic phenotypes [8]. Knowing that most genes require a network of other genes to produce a biological effect, mastery of gene regulation now plays a crucial role in elucidating the functional context of genes within the genome that involves complex biological processes [9]. The aim of this review was to identify a network of genes whose expression is associated with KLF14 variation in trans and the protein-protein interactions in the KLF14 protein network, providing a framework for understanding how KLF14 could influence disease risk.
Material and Methods
Data sources
Putative transcription factors within 10-20 kb upstream and 10 kb downstream of KLF14 gene were searched with data extraction from SABiosciences web-base (https://www. sabiosciences.com/) of the QIAGEN company (https:// www.qiagen.com/ca/). Specifically for Human (Homo sapiens), data on transcriptions factors were extracted from GenBank of the National Centre for Biological Information (https:// www.ncbi.nlm.nih.gov/). The names, chromosomal locations, gene IDs, previous symbols and aliases, characteristics and functions for these genes are given on the NCBI website (https://www.ncbi.nlm.nih.gov/genome/guide/human/). The online software STRING v9.1 was used to identify the known and predicted protein-protein interaction in the KLF14 protein network. STRING is a comprehensive, authoritative database currently covers more than 9,643,763 proteins from 2031 organisms and, mainly focusing on predicted and known protein interactions. These interactions include direct (physical) and indirect (functional) associations.
Construction of genes-associated transregulatory network and protein-protein interactions network
The genes associated in trans regulation in the transcription of KLF14 gene were positioned according to their chromosomal locations in the human karyotype given by NCBI website. The protein-protein interactions of KLF14 protein network was built with STRING online software v9.1 under the active prediction methods (Gene Fusion, Neighborhood, Cooccurrence, Co-expression, Experiments, Databases and Text Mining), Medium Confidence (0.400) and no more than 50 interactions parameters.
Results
Genes associated trans-regulatory network
Through the NCBI database, from 82 (eighty-two) transcription factors implicated in trans-regulation of KLF14 gene identified, only 62 (sixty two) was present in human (Homo sapiens) (Figure 1). The genes that encode for these transcriptions factors plays roles in a wide variety of processes including: fetal development (PAX3); hematopoiesis, apoptosis, development and cell differentiation and proliferation (EVI1); control of pattern formation during development of the central nervous system (EN1); crucial in normal development (MEIS1) development and proliferation of hematopoietic and endocrine cell lineage (GATA2); important regulator in early development (PAX5); development of the eye, tooth and abdominal organs (PITX2); ocular development (FOXC1); muscle development (EGR3, MEF2A); lymphocyte development, endothelial cell growth and migration (EGR3); neuronal development (EGR3, PAX5 ); erythroid development (GATA1); development of the central nervous system (EN1); cell cycle (E2F3, E2F4); cell growth control (MEF2A); endothelial cell (GATA3, EGR3); terminal differentiation of somatotroph and lactotroph cell phenotypes (PITX2); muscle regeneration (MYOD1); hematopoiesis (EVI1); apoptosis (EVI1, MEF2A); differentiation and proliferation (EVI1); immunoregulation and inflammation (IL10); tumorigenesis (cmyb); osteogenesis (PBX1); spermatogenesis (PAX5); regulation of embryonic (FOXC1); neuronal differentiation (MEF2A); formation of blood vessels (ANGPTL1).
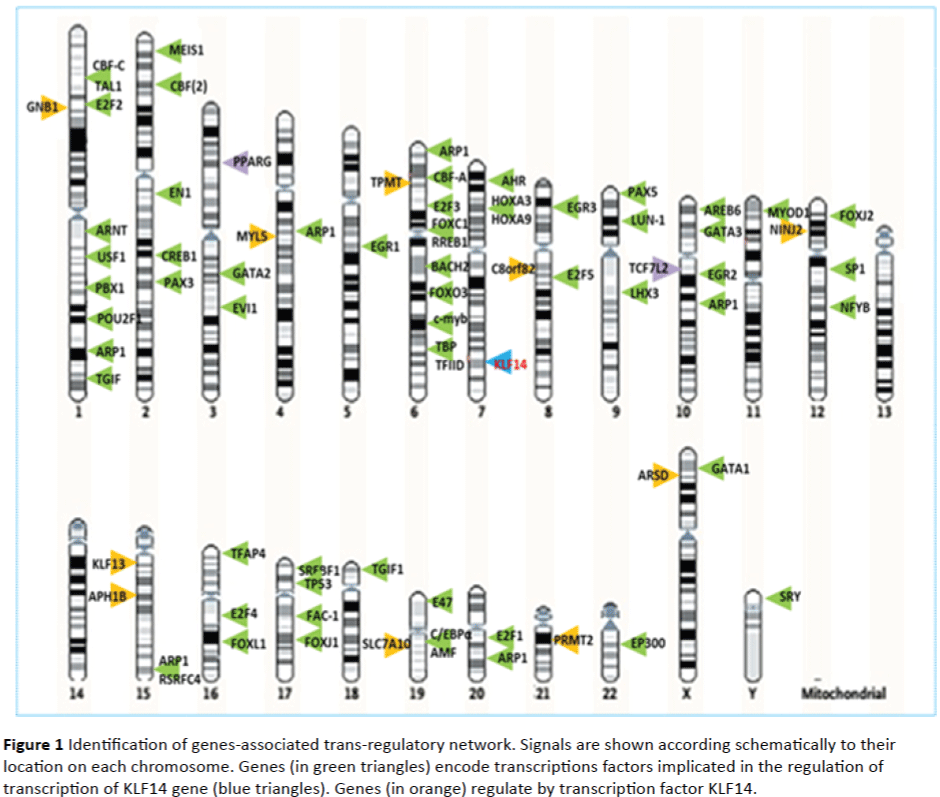
Figure 1: Identification of genes-associated trans-regulatory network. Signals are shown according schematically to their location on each chromosome. Genes (in green triangles) encode transcriptions factors implicated in the regulation of transcription of KLF14 gene (blue triangles). Genes (in orange) regulate by transcription factor KLF14.
Interestingly, TCF7L2 and PPARG gene which are the main genes implicated in the pathophysiology of metabolic diseases are not present among genes which encode putative transcriptions factors involved in the trans-regulation of KLF14 gene. The KLF14 gene encodes for a transcription factor which acts in trans to regulate the expression of a network of genes associated with metabolic traits, (body mass index, glucose levels, high density lipoprotein levels, index of insulin sensitivity, insulin levels, low density lipoprotein levels, triglyceride levels, T2DM, waist-hip ratio). This transcription factor also interact with other proteins which expression are implicated in many others physiological processes.
Protein-protein interactions network
Some 32 proteins were identified (ZFAND6, SIN3A, ARAP1, XRN1, ZBED3, SLC30A8, CHST8, KDM2A, FBXL19, KDM2B, MTNR1B, CDKAL1, ADCY5, PTPN23, RANBP2, KCNQ1, C2CD4B, IGF2BP2, TTC39B, CAMK1D, TCF7L2, JAZF1, CAPN13, TP53INP1, AGMO, KDM6A, UTY, KDM6B, C6orf106, UBC, FTO and CDC123) to interact with KLF14 protein resulting in a network diagram with 33 nodes (gene/proteins) including the KLF14 node and with 50 direct edges or interactions (Figure 2). By these interactions, KLF14 protein could influence the action of these 32 proteins and the processes in which they are implicated. It is important here to note the presence in this diagram of other gene/proteins previously shown to be associated with obesity such as the (FTO, TCF7L2) [7]; with T2DM (ZFAND6, SLC30A8, MTNR1B, CDKAL1, KCNQ1, IGF2BP2, TCF7L2, JAZF1, MTNR1B, FTO, CDC123) [7]. This network also shows that the KLF14 protein interacts with UBC, a polyubiquitin precursor, a member of ubiquitin family with its highly conserved 76 amino acid proteins. They are abundant (0.1%-5% of total proteins) in eukaryotic cells, and can be conjugated to other proteins via an isopeptide linkage through a carboxy-terminal glycine residue of ubiquitin and the ε- amino group of a lysine in the target protein. They can equally make an isopeptide bond with a lysine in another moiety of ubiquitin to form UB chains. This ubiquitination of a target protein has been associated with protein degradation through the 26S proteasome, protein trafficking, kinase modification, endocytosis, cell cycle regulation, DNA repair, apoptosis, and regulation of other cell signaling pathways [10-12]. The amount of ubiquitin protein in a cell is maintained at an adequate level depending on cell conditions, despite its use and large number of substrates which are ubiquitinated [13,14], this homeostasis is established by recycling of ubiquitin chains by deubiquitinating enzymes and de novo synthesis [15].
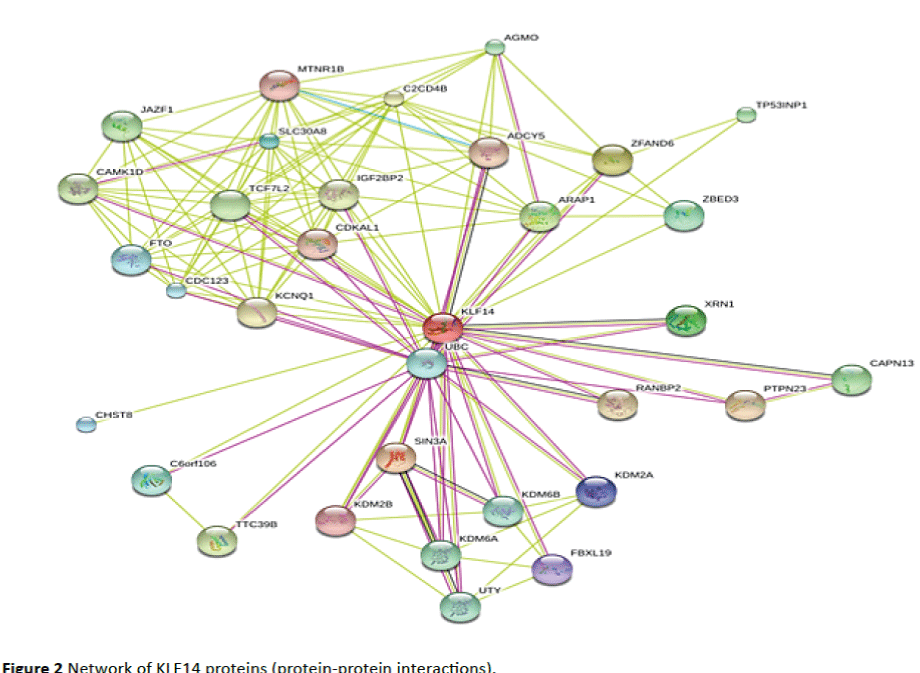
Figure 2: Network of KLF14 proteins (protein-protein interactions).
Putative regulatory network of KLF14 (gene/ protein)
Upstream of KLF14 gene located on 7q32.3 chromosome, there are sixty two (62) transcriptions factors (with three of them encode by AHR, HOXA3 and HOXA9 genes located on the same chromosome) that came to bind within 10-20 Kb upstream and 10 Kb downstream of KLF14 gene for his transcription. This transcription is repressed by Sin3A protein which leads to the HDAC1 and HDAC2 enzymatic proteins important for transcription repression, Mad and MeCP2 (DNA binding proteins) and the Ikaros and SMRT (co-repressor) [16,17]. At the end of the KLF14 gene transcription, the resulting mRNA is translated in KLF14 protein. This protein (transcription factor) act on ten genes identified to have GWST association driven by rs4731702 (C/T) of KLF14 gene and established protein-protein interactions with 32 proteins implicated in metabolic diseases and others biological processes. Of particular are the presence UBC and TCF7L2 (highly expressed in most human tissues and which influences insulin sensitivity in skeletal muscle via indirect pathways, insulin resistance through altered endocrine function of adipocytes and reduced β-cell function [18]) (Figure 3). The physical interactions which exist between KLF14 and UBC could regulate the amount of KLF14 protein available by its degradation.
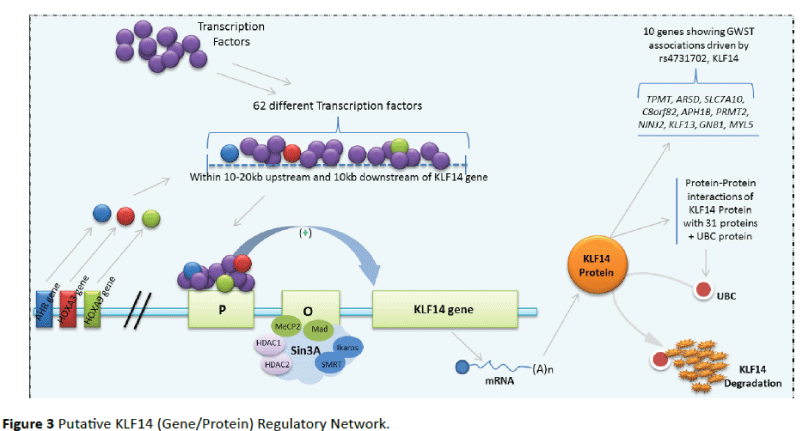
Figure 3: Putative KLF14 (Gene/Protein) Regulatory Network.
Conclusion
Knowledge of genetic information and the mastery of the vast network of regulation of the KLF14 (Gene/Protein) and others genes implicated in metabolic diseases such as T2DM and Obesity could be used as a source of information for new and existing targets to interactions and characterize high-risk populations point towards a trustworthy direction for clinical or fundamental research. This provides a variety drug development targets in the form of signal pathways that can reduce the progression of multiple metabolic phenotypes or the risk of their complications.
Author Contribution
MGF: Designed the study and acquired data; analyzed data; interpreted data; drafted and revised the manuscript for intellectual content. BAT: Revised the manuscript for intellectual content. ES: Revised the manuscript for intellectual content. JCM: Revised the manuscript for intellectual content. WFM: Conceived and designed the study; analyzed data; interpreted data; drafted the manuscript; supervised the study and revised the manuscript for intellectual content.
All authors had full access to the data and approved the final version of the manuscript.
Conflict of Interest
None
Acknowledgements
We gratefully acknowledge the Laboratory of Public Health Research Biotechnology (LAPHER-BIOTECH) and the Laboratory of Molecular Medicine and Metabolism (LMMM) at the Biotechnology Center of the University of Yaoundé I.
9555
References
- Van Dieren S, Beulens JW, van der Schouw YT, Grobbee DE, Neal B (2010) The global burden of diabetes and its complications: an emerging pandemic. Eur J Cardiovasc Prev Rehabil 17: 3-8.
- International Diabetes Federation: IDF Diabetes Atlas (2015) (7thedn). Available at: www.diabetesatlas.org.
- Zimmet P, Alberti KG, Shaw J (2001) Global and societal implications of the diabetes epidemic. Nature414: 782-787.
- Waterfield T, Gloyn AL (2008) Monogenic β-cell dysfunction in children: clinical phenotypes, genetic etiology and mutational pathways. Pediatr Health 2: 517-532.
- Hattersley AT, McCarthy MI (2005) What makes a good genetic association study? Lancet 366: 1315-1323.
- Sladek R, Rocheleau G, Rung J, Dina C, Shen L, et al. (2007) A genome-wide association study identifies novel risk loci for type 2 diabetes. Nature 445:881-885.
- McCarthy MI (2010) Genomics, Type 2 Diabetes, and Obesity. New England Journal of Medicine 363: 2339-2350.
- Kerrin SS, Asa KH, Grundberg E, Nica AC, Thorleifsson G, et al. (2011) Identification of an imprinted master trans-regulator at the KLF14 locus related to multiple metabolic phenotypes. Nature Genetic43: 561-564.
- Pickart CM (2001) Mechanisms underlying ubiquitination. Annual Review of Biochemistry 70: 503-533.
- Mukhopadhyay D, Riezman H (2007) Proteasomeindependent functions of ubiquitin in endocytosis and signaling. Science 315: 201-205.
- Clague MJ, Urbé S (2010) Ubiquitin: Same molecule, different degradation pathways. Cell 143: 682-685.
- Ryu KY, Baker RT, Kopito RR (2006) Ubiquitin-specific protease 2 as a tool for quantification of total ubiquitin levels in biological specimens. Analytical Biochemistry 353: 153-155.
- Crinelli R, Bianchi M, Menotta M, Carloni E, Giacomini E, et al. (2008) Ubiquitin over-expression promotes E6AP autodegradation and reactivation of the p53/MDM2 pathway in HeLa cells. Molecular and Cellular Biochemistry 318: 129-145.
- Kimura Y, Tanaka K (2010) Regulatory mechanisms involved in the control of ubiquitin homeostasis. The Journal of Biochemistry 147: 793-798.
- Cunliffe VT (2008) Eloquent silence: developmental functions of Class I histone deacetylases. Current Opinion in Genetics and Development 18: 404-410.
- Kadosh D, Struhl K (1998) Histone deacetylase activity of Rpd3 is important for transcriptional repression in vivo. Genes and Development 12: 797-805.
- Cauchi S, Meyre D, Dina C, Choquet H, Samson C, et al. (2006) Transcription factor TCF7L2 genetic study in the French population: expression in human beta-cells and adipose tissue and strong association with type 2 diabetes. Diabetes 55: 2903-2908.