Keywords
Fish waste recovery; Detergent proteases; Sustainable detergents; Enzyme biotechnology; Serine proteases
Introduction
The mutton snapper (Lutjanus analis) is distributed throughout the Atlantic west coast, from North Carolina (USA) to southeastern Brazil. These species forms large shoals, usually on hard substrates at depths between 10 and 100 m. Juveniles live in clusters in coastal waters, typically associated with reefs (Allen GR 1985, Manooch CSIII and Drennon CL 1987). From a commercial standpoint, mutton snapper is very important for small local fisheries and markets, having contributed to 0.62% (3,014.5 tons) of total Brazilian marine capture production in 2011 (MPA, 2013). The mutton snapper and other fish must undergo evisceration and/ or fileting after they are captured. These processes add value to the product; however, they also generate several organic wastes such as bones, viscera, skins, and carcasses. Such residues can be harmful to the environment, even in small amounts. Nevertheless, they are rich in bioactive molecules and can be used to obtain fish meal and oil, collagen, gelatin, leather, and enzymes (Blanco M et al., 2007). Among these products, enzymes comprise a group with a high market value, which reached $4.8 billion in 2013 (BCC Research Report Highlights: https://www.bccresearch.com/ market-research/biotechnology/enzymes-industrial-applicationsbio030h. html “Accessed 12 Feb 2015”). Technical enzymes are the second most valuable category with the greatest volume being used in the detergent industry (Aberer W et al., 2002). The main enzymes used in detergents are proteases. Otto Röhm first proposed this concept by introducing pancreatic extract in washing powder composition in 1913 (Maurer K-H 2004). Nowadays, subtilisin (from Baccilus licheniformis) has replaced mammal digestive enzymes as a laundry detergent additive (Otzen D, 2011).
Even though enzymes from mammals are not considered to be efficient for use in laundry detergents, digestive enzymes from other animals are suitable for such applications. Different from their mammalian counterparts, fish enzymes have higher catalytic efficiency and stability, and therefore present great potential for numerous commercial uses (Jónsdóttir G et al., 2004). Enzymes that are more specifically targeted for use as laundry detergent additives must be stable at high pH and temperatures and be compatible with different types of surfactants and oxidizing agents. These characteristics have been found in fish enzymes, such as trypsin from zebra blenny (Salaria basilisca) (Ktari N et al., 2012). Alkaline proteases from grey triggerfish (Balistes capriscus) (Jellouli K et al., 2009), and tambaqui (Colossoma macropomum) (Espósito TS et al., 2009).
In this paper, we present the partial purification process of alkaline proteolytic enzymes from mutton snapper and provide information about their biochemical characteristics as well as their compatibility with commercial laundry detergents, oxidants, and surfactant agents.
Materials and Methods
Enzyme extraction
Adult specimens of mutton snapper (L. analis) were obtained from fishermen of Pina beach on the coast of Recife, Pernambuco, Brazil. The animals (n=3) presented an average weight of 866.7 ± 110.0 g and length of 31.3 ± 3.05 cm. The intestines (7.86 g) were collected and immediately homogenized in 0.1 M Tris-HCl and 15 mM NaCl (pH 8.0) at 4°C (100 mg/mL). The resulting homogenate was centrifuged (MR Herolab UniCen, Germany) at 10,000 xg for 15 min at 4°C to remove cell debris. The supernatant (crude extract) was maintained at −20°C for subsequent purification steps (Bezerra RS, et al., 2005).
Enzyme partial purification
For the partial purification of the enzymes, 78 mL of the intestine crude extract was first heat treated at 45°C for 30 min and centrifuged at 10,000 xg for 15 min at 4°C. The supernatant was collected and fractioned with ammonium sulfate to obtain a final salt concentration of 80%. After 2 h at 4°C, the sample was centrifuged at 10,000 xg for 15 min at 4°C. The precipitate (fraction 0%-80%) was resuspended in 10 mL of 0.1 M Tris-HCl buffer (pH 8.0) and dialyzed against this same buffer for 24 h at 4°C [14]. Thereafter, approximately 4.2 mL (6.0 mg protein) of the 0%-80% fraction was added into a Sephadex™ G-75 gel filtration column at a flow rate of 0.3 mL/min. The chromatography fractions were checked for the presence of protein (absorbance at 280 nm) and trypsin activity. Fractions with higher trypsin activity were pooled and stored at −20°C for subsequent experiments.
Trypsin activity
The tryptic activity of the samples was estimated using an 8 mM benzoyl-arginine-p-nitroanilide (BApNA) substrate prepared in dimethyl sulfoxide (DMSO). For the assay, 30 μL of the sample was mixed with 140 μL of 0.1 M Tris-HCl buffer, pH 8.0, and 30 μL of 8 mM BApNA for 15 min at 25°C. The formation of p-nitroaniline (product) was measured with a microplate reader (Bio-Rad xMark spectrophotometer, California, USA) at 405 nm. A blank control, in which the sample was replaced by distilled water, was prepared. One unit of enzyme activity was defined as the amount of enzyme required to hydrolyze 1 μmol/min of BApNA (Erlanger BF, 1961). The total protein content of the samples was estimated using the procedure described by Bradford (Bradford MM, 1976) with bovine serum albumin (BSA) as the standard protein.
Total protease activity
Alkaline protease activity was assayed using 1% (w/v) azocasein (Sigma-Aldrich®) substrate prepared in 0.1 M Tris-HCl, pH 7.2 (Bezerra RS et al., 2005, Alencar RB et al., 2003). The enzymatic reaction was prepared in triplicate using microcentrifuge tubes to which 50 μL of 1% (w/v) azocasein and 30 μL of sample were added. After 60 min at 25°C, 240 μL of 10% (w/v) trichloroacetic acid was added to stop the reaction, and the mixture was centrifuged at 8,000 xg for 5 min. Subsequently, 70 μL of supernatant was added to 130 μL of 1 M NaOH, and the absorbance of the mixture was measured with a microplate reader (according to the above item) at 450 nm. A blank control, in which the sample was replaced by distilled water, was prepared. One unit (U) of enzyme activity was defined as the amount of enzyme needed to hydrolyze azocasein to produce an absorbance change of 0.001/min.
Sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE)
SDS-PAGE was performed using the procedure described by Laemmli UK (1970) with a 4% (w/v) stacking gel and a 12.5% (w/v) separation gel. The gel was stained for protein identification with 0.01% (w/v) Coomassie Brilliant Blue R-250 dissolved in 10% (v/v) acetic acid and 25% (v/v) methyl alcohol. The backside of the gel was destained by washing with the same solution without Coomassie.
Effect of protease inhibitors
This assay was performed with the following synthetic protease inhibitors: phenylmethylsulfonylfluoride (PMSF, serine protease inhibitor), tosyl lysine chloromethyl ketone (TLCK, trypsin inhibitor), benzamidine (trypsin inhibitor), and tosyl phenylalanine chloromethyl ketone (TPCK, chymotrypsin inhibitor) at 8 mM dissolved in DMSO and ethylenediaminetetraacetic acid (EDTA, metalloprotease inhibitor) and β-mercaptoethanol (reducing agent) at 8 mM prepared in water. The inhibitors were incubated with the partially purified enzyme in a proportion of 1:1 (v/v) for 30 min at 25°C. Thereafter, 30 μL of incubate was mixed with 50 μL of 1% (w/v) azocasein for 60 min at 25°C. The remaining steps of the assay were performed as previously described in the total protease activity section. The residual activity of each sample was estimated by comparing it with a positive control (100% activity), in which the sample was incubated with DMSO or water depending on the solvent of the inhibitor (Bezerra RS et al., 2001)
Effect of pH and temperature
The influence of pH on the proteolytic activity of the partially purified enzyme was analyzed with different 0.2 M buffer solutions (citrate phosphate, pH 2.0-7.5; Tris-HCl, pH 7.0-9.0; and NaOHglycine, pH 8.7-12.0) at 25°C for 15 min using BApNA as the substrate. The assays of temperature were performed in a water bath (Bezerra RS et al., 2005) at temperatures ranging from 25 to 75°C. The influence of temperature on the proteolytic activity was investigated by incubating the enzyme with 8 mM BApNA for 15 min at each temperature. For the thermal stability test, the partially purified proteases were incubated for 15 min at different temperatures and then incubated with 8 mM BApNA for 15 min at 25°C. The obtained products were measured in triplicate at 405 nm.
Trypsin kinetics
The kinetic parameters (Vmax and Km) of the partially purified proteases were determined with BApNA as substrate. The final substrate concentration ranged from 0.04 to 4.80 mM. The asymptotic parameters were calculated with the Origin software program, version 6.0 (Microcal Software, Inc.).
Effect of oxidizing agents and surfactants
The stability of the partially purified sample was tested in the presence of hydrogen peroxide (5%, 10%, and 15% v/v) and 1% solutions of different anionic (sodium dodecyl sulfate-SDS and sodium cholate), zwitterionic (CHAPS), and nonionic (Triton X-100, Tween 80, and Tween 20) surfactants and a commercial detergent (Extran®). The assays were performed by incubating the partially purified enzyme with the oxidizing agents or surfactant in a ratio of 1:1 (v/v) for 30 and 60 min at 25°C (Moreira KA, 2002) Subsequently, 30 μL of the incubated solution was mixed with 50 μL of 1% (w/v) azocasein for 60 min at 25°C. A blank assay (negative control) was performed by replacing the enzyme with the buffer in the reaction. The remaining steps of the assay were performed as previously described in the total protease activity section. The residual activity of each sample was estimated by comparing it with a positive control (100% activity), in which the enzyme sample was incubated with water.
Compatibility with commercial detergents
The partially purified enzyme from mutton snapper was incubated at 25°C with the following commercial detergents: Ala® and Ace® (Procter & Gamble), Bem-te-vi® (Alimonda), Surf® (UniLever), and Assim® (Assolan) at a final concentration of 7 mg/mL. The samples were agitated and collected at intervals of 30 and 60 min at 25°C. Thereafter, 30 μL of the incubated sample was mixed with 50 μL of 1% (w/v) azocasein for 60 min at 25°C. The remaining steps of the assay were performed as previously described in the total protease activity section. The residual activity of each sample was estimated by comparing it with a positive control (100% activity), in which the sample was incubated with water and without commercial detergents. The residual proteolytic activity of each sample was compared with a control sample of detergent incubated with Tris-HCl, pH 8.0, at 25°C (Moreira KA, 2002).
Results
The three-step partial purification process of the alkaline proteases from the intestine of mutton snapper (L. analis) is summarized in Table 1. After the first step (heat treatment), the specific activity was 1.1 times higher than initial crude extract activity (0.715 U/mg), and the yield was 101%. The second (ammonium sulfate fractionation) and third (Sephadex™ G-75 gel filtration column) steps presented yields of 31% and 10%, respectively. The results of the SDS-PAGE (Figure 1) show proteins between 55 and 14 kDa.
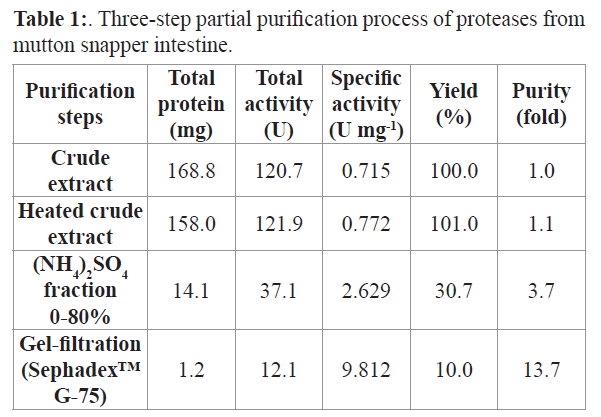
Table 1:. Three-step partial purification process of proteases from mutton snapper intestine.
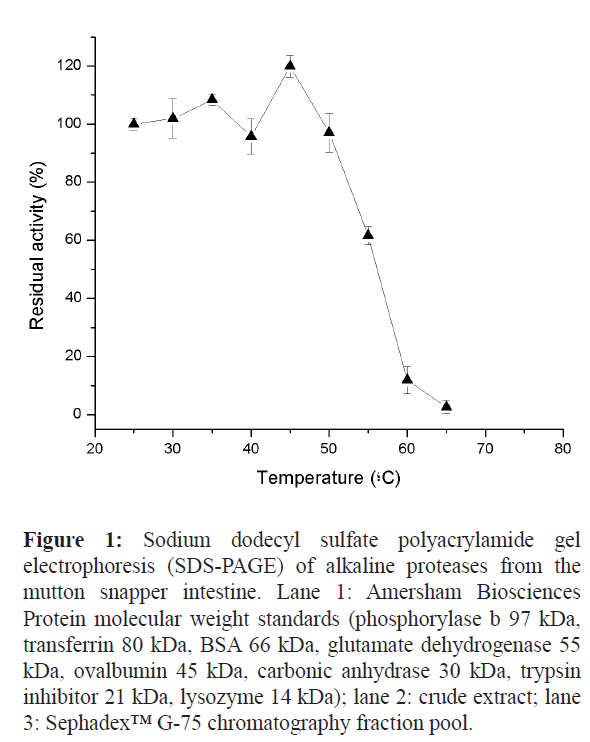
Figure 1: Sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) of alkaline proteases from the mutton snapper intestine. Lane 1: Amersham Biosciences Protein molecular weight standards (phosphorylase b 97 kDa, transferrin 80 kDa, BSA 66 kDa, glutamate dehydrogenase 55 kDa, ovalbumin 45 kDa, carbonic anhydrase 30 kDa, trypsin inhibitor 21 kDa, lysozyme 14 kDa); lane 2: crude extract; lane 3: Sephadex™ G-75 chromatography fraction pool.
The proteolytic activity (with azocasein as substrate) of the partially purified proteases from mutton snapper was strongly inhibited by PMSF (60.7%), TLCK (77.9%), and benzamidine (63.7%) (Table 2). The classic inhibitor for chymotrypsin TPCK inhibited approximately 30% of the activity. Likewise, β-mercaptoethanol did not produce any inhibitory effect on the proteases, indicating that the reduction of their disulfide bonds did not affect their activity
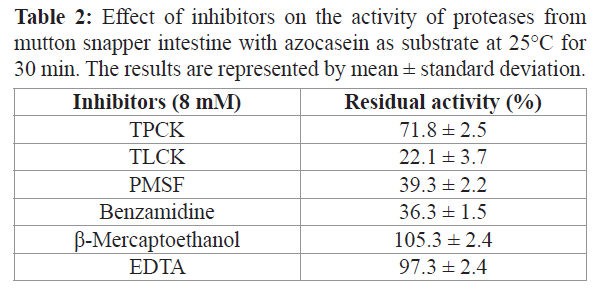
Table 2: Effect of inhibitors on the activity of proteases from mutton snapper intestine with azocasein as substrate at 25°C for 30 min. The results are represented by mean ± standard deviation.
The purified proteases have a Km value of 0.564 mM and Vmax value of 0.609 U, with BApNA as substrate. This Km value is similar to those obtained for lane snapper (Lutjanus synagris) trypsin (Espósito TS, 2010) and anchovy (Engraulis encrasicolus) trypsin (Martínez A, 1988) However, these proteases exhibited higher affinity to BApNA than bovine (0.939 mM) and hybrid tilapia trypsins (Bezerra RS, 2005, Erlanger BF, 1961). With the BApNA substrate, the partially purified proteases from the intestine of mutton snapper were most active in a pH range of 8.0- 11.5. The purifyed proteases were very active within a temperature range of 35°C-60°C. A higher activity of this enzyme was found at 55°C similar to tambaqui (Bezerra RS, 2001) and carp (Martínez A, 1988) trypsins. As regards thermal stability, the mutton snapper proteases maintained up to 60% of their activity after 15 min at 55°C.
In regard to detergent components, the proteases from mutton snapper were not inhibited or activated after 60 min of incubation with hydrogen peroxide (H2O2) at concentrations ranging from 5% to 15% in increments of 5% (Table 3). The porcine trypsin, on the other hand, was significantly activated in the presence of these hydrogen peroxide concentrations. Another important component of detergents is surfactant agents. The nonionic surfactants Triton X-100, Tween 20, and Tween 80 activated the mutton snapper enzyme to approximately 60% whereas the porcine trypsin was not inhibited or activated by them. The zwitterionic surfactant CHAPS activated the mutton snapper enzymes to 17.1% after 30 min; however, after 60 min, it inhibited the enzymes by 17.2%. Both porcine trypsin and mutton snapper were inhibited by anionic surfactants (SDS and sodium cholate) after 30 min of incubation (Table 3). However, the activity of mutton snapper proteases increased after 60 min of incubation with sodium cholate.
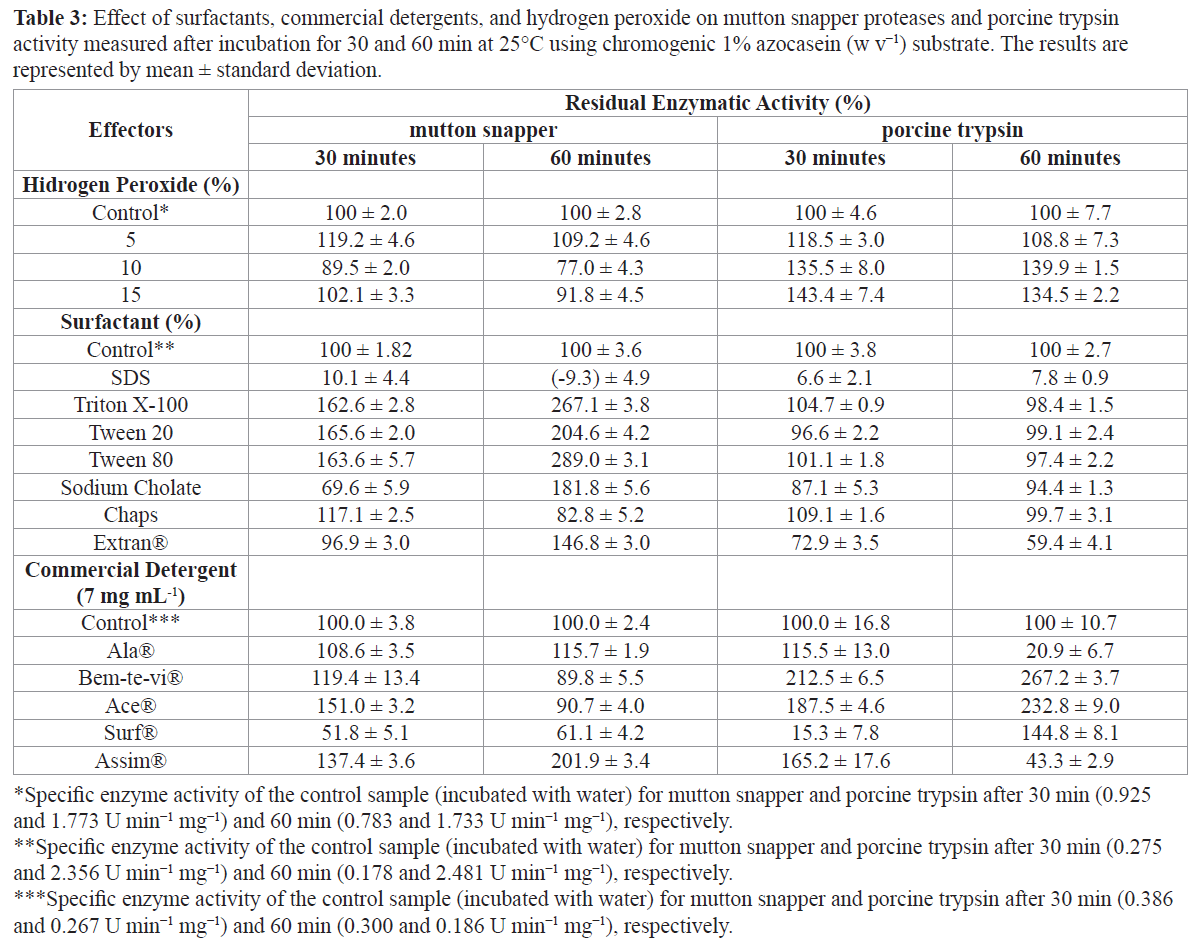
Table 3: Effect of surfactants, commercial detergents, and hydrogen peroxide on mutton snapper proteases and porcine trypsin activity measured after incubation for 30 and 60 min at 25°C using chromogenic 1% azocasein (w v−1) substrate. The results are represented by mean ± standard deviation.
The alkaline proteases from mutton snapper were stable and compatible with most commercial detergents (Table 3). The enzyme was activated in the presence of Ala®, Bem-te-vi®, Ace®, and Assim® (8.6%, 19.4%, 51.1%, and 37.4% activation, respectively) after 30 min of incubation and inhibited in the presence of Surf® (48.2% inhibition). Maximum stability was observed with Assim® after 60 min, in which the enzymes were activated to 101.9%. Porcine trypsin maintained more than 100 % of its activity when incubated for 30 min with all commercial detergents. However, after 60 min, Ala® and Assim® inhibited the porcine trypsin to 79.1% and 56.7%, respectively (Table 3).
Discussion
Even though a single purified enzyme was not obtained in the present work, for the current application (laundry detergent), a mixture of proteases rather than purified preparations are useful because a wider range of substrates must be hydrolyzed. The inhibition results indicate that the sample is composed primarily of serine proteases, particularly trypsin and chymotrypsin-like enzymes, which are the main proteases found in fish digestive tracts. Similar results were also observed for other tropical fish proteases [Jellouli K, 2009, Bezerra RS, 2005, Ali NEH, 2009, Marcuschi M et al., 2010). The lack of inhibition in the presence of EDTA (metalloenzyme inhibitor) indicates that the activity of these proteases is not ion dependent. Chelating agents are commonly found in powder detergent formulations and act as water softeners and also assist in stain removal (Ktari N, 2010). Therefore, enzymes must be stable to these agents as well.
Studies show that the activity of serine proteases can be approximately 70% lower at pH 6.0 because of the protonation from the histidine in the active site, thus significantly reducing enzyme activity [24]. On the other hand, a high pH value can also decrease catalytic activity (Kasserra HP, 1969). However, in the present study, the catalytic activity of the enzyme was not reduced with increasing pH, which may be attributed to the high affinity of the substrate to the enzyme. Similar results were observed for other fish when the same substrate was used (Ktari N, 2012, Kurtovic I, 2006, Klomklao S, 2007). Enzymes that are highly active in alkaline pH are usually targeted for application in laundry detergents because their pH typically ranges between 9.0 and 12.0.
Another important characteristic sought in enzymes targeted for laundry detergents is thermal stability between 50 and 60°C and high activity in a wide range of temperatures. These characteristics are considered to satisfy both the “hot wash” [Jellouli K, 2009, Moreira KA, 2002, Espósito TS, 2009) and “cold wash,” which is more environmentally friendly, features found in many washing machines. The stability at higher temperatures observed the for mutton snapper enzyme may be related to the temperature selection in the heat treatment step during the purification process.
Proteases used in detergent formulations must also be stable in the presence of different soap components. The stability of the mutton snapper proteases was compared with that of a commercial porcine trypsin (Sigma-Aldrich®) in the presence of oxidants, surfactants, and detergents at 25°C. Typically, oxidizing agents may denature proteins, primarily by oxidizing their methionine residues, thus facilitating the removal of proteinaceous stains (Maurer K-H, 2004) However, this may also pose a problem for the enzymes present in detergents because they may lose their activity in the presence of these components.
Activation of trypsin-like enzymes in these concentrations was also seen for common carp (Cyprinus carpio) (Espósito TS, 2009) The activation of trypsin by hydrogen peroxide is not yet well understood; however, these enzymes are probably not inactivated because their activity is not dependent on cysteine or methionine residues.
According to Malhotra and Coupland (Malhotra A, 2004) nonionic surfactants, such as Triton X-100, Tween 20, and Tween 80, cannot bind to the surface of globular proteins. Therefore, nonionic surfactants typically do not affect protein conformation or induce association with other proteins. Moreover, nonionic surfactants can also shield the hydrophobic core of proteins from the solvent, thus increasing its solubility. On the other hand, SDS is a charged surfactant; therefore, it can interact with the charged surface and hydrophobic core of proteins. Therefore, charged surfactants can act as denaturing agents by affecting protein solubility and conformation through the formation of small aggregates around their polypeptide backbone (Malhotra A, 2004, Ghosh S, 2008).
These results regarding stability to powder detergents clearly indicate that the performance of enzymes in detergents depends on a number of factors, including the compounds of detergent because the proteolytic activity varied with each laundry detergent (Ktari N, 2012).
Acknowledgements
This study was financially supported by the following Brazilian agencies: Ministry of Fisheries and Aquaculture, Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES), Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq), Financiadora de Estudos e Projetos (FINEP), Fundação de Amparo à Ciência e Tecnologia de Pernambuco (FACEPE) and PETROBRAS.
6892
References
- Allen,G.R.,(1985).FAO Species Catalogue, Fish Synop (125) 6 , FAO, Rome.
- nManooch, C.SIIIDrennon,C.L., (1987).Age and growth of yellowtail snapper and queen triggerfish collected from the U.S. Virgin Islands and Puerto Rico Fish, Res, 6 (1):53-68.
- nMPA.,(2013). Statistical Bulletin of Fisheries and Aquaculture, 2011 Brasília: Fisheries and AquacultureMinistry (In Portuguese).
- nBorderías, A.J. Sánchez-Alonso, I., (2011).First Processing Steps and the Quality of Wild and Farmed Fish J Food Sci 76 (1):1-5.
- nBlanco, M. Sotelo, C.G. Chapela, M.J. Pérez-Martín, R.I., (2007). Fish species identification in seafood products Trends Food Sci Tech 18 :29-36.
- nAberer, W. Hahn, M. Klade, M.Seebacher, U.Spök, A. Wallner, K.Witzani, H., (2002).Collection of information on enzymes Office for Official Publications of the European Communities, Luxembourg
- nMaurer, K-H., (2004). Detergent peptidasesCurrOpinBiotechnol 15 :330-334.
- nOtzen, D., (2011). Protein-surfactant interactions: A tale of many states BiochimicaetBiophysicaActa 1814 :562-591
- nJónsdóttir, G., Bjarnason, J.B., Gudmundsdóttir. A., (2004). Recombinant cold-adapted trypsin I from Atlantic cod—expression, purification, and identification Protein Express Purif 33 :110-122
- nKtari, N., Khaled, H.B., Nasri, R., Jellouli, K., Ghorbel, S., Nasri, M., (2012). Trypsin from zebra blenny (Salariabasilisca) viscera: Purification, characterisation and potential application as a detergent additive Food Chem 130: 467-474
- nJellouli, K.,Bougatef, A., Daassi, D., Balti, R.,Barkia, A., Nasri, M., (2009). New alkaline trypsin from the intestine of Grey triggerfish (Balistescapriscus) with high activity at low temperature: Purification and characterization Food Chem 116 :644-650
- nEspósito, T.S.,Amaral, I.P.G.,Buarque, D.S., Oliveira, G.B.,Carvalho, Jr.L.B.,Bezerra, R.S., (2009). Fish processing waste as a source of alkaline proteases for laundry detergent Food Chem 112 :125-130
- nBezerra, R.S., Lins, E.J.F., Alencar,R.B., aiva, M.G., Chaves,M.E.C., Coelho,L.C.B.B., and Carvalho, jr.L.B., (2005). Purification and Characterization of a Protease Produced by a Planomicrobiumsp L-2 from Gut of Octopus vulgaris Process Biochem 40 :1829-1834
- nBezerra, R.S. Santos, J.F. Paiva, P.M.G.Correia,M.T.S. Coelho, L.C.B.B. Vieira, V.L.A. Carvalhojr.L.B., (2001). Partial purification and characterization of a thermostable trypsin from pyloric caeca of tambaqui (Colossomamacropomum) J Food Biochem 25: 199-210
- nErlanger, B.F. Kokowsky, N. Cohen, W., (1961). The Preparation and Properties of Two New Chromogenic Substrates of Trypsin Arch BiochemBiophys 95: 271-278
- nBradford, M.M., (1976). A Rapid and Sensitive Method for the Quantitation of Microgram Quantities of Protein Utilizing the Principle of Protein-Dye Binding Anal Biochem 72 :248-254
- nAlencar, R.B. Biondi,M.M. Paiva, P.M.G. Vieira, V.L.A.CarvalhoJr, L.B. Bezerra, R.S. Alkaline Proteases from the Digestive Tract of Four Tropical Fishes (2003).Braz J Food Technol 6: 279-284
- nLaemmli, U.K., (1970). Cleavage of Structural Proteins during the Assembly of the Head of Bacteriophage T4 Nature 227: 680-685
- nMoreira, K.A. Albuquerque, B.F. Teixeira, M.F.S. Porto, A.L.F, Lima Filho, J.L., (2002). Application of protease from Nocardiopsissp as a laundry detergent additive World J MicrobiolBiotechnol 18 :307-312
- nEspósito, T.S. Marcuschi, M. Amara,I.P.G. Carvalho, Jr.L.B. Bezerra, R.S., (2010). Trypsin from the Processing Waste of the Lane Snapper (Lutjanussynagris) and Its Compatibility with Oxidants, Surfactants and Commercial Detergents J Agric Food Chem 58 :6433-6439
- nMartínez, A. Olsen, R.L. Serra,J.L., (1988). Purification and characterization of two trypsin-like enzymes from the digestive tract of anchovy engraulisencrasicholus Comp BiochemPhysiol B 91 :677- 684
- nEspósito, T.S.Amaral, I.P.G. Marcuschi, CarvalhoJrL.B. Bezerra, R.S., (2009). Surfactants- and oxidants-resistant alkaline proteases from common carp (Cyprinuscarpio l) processing waste J Food Biochem 33: 821-834
- nAli, N.E.H.Hmidet, N.Bougatef, A.Nasri, R.Nasri, M., (2009). A Laundry Detergent-Stable Alkaline Trypsin from Striped Seabream (Lithognathusmormyrus) Viscera: Purification and Characterization J Agric Food Chem 57: 10943-10950
- nMarcuschi, M.Espósito, T.S. Machado, M.F.M. Hirata, I.Y. Machado, M.F.M. Silva, M.V.Carvalho, Jr.L.B. Oliveira, V. Bezerra, R.S., (2010). Purification, characterization and substrate specificity of a trypsin from the Amazonian fish tambaqui (Colossomamacropomum) BiochemBioph Res Co 396: 667-673
- nKasserra, H.P.Laidler, K.J., (1969). Mechanisms of action of trypsin and chymotrypsin Can J Chem 47 :4031-4039
- nKurtovic I. Marshall, S.N. Simpson, B.K., (2006). Isolation and characterization of a trypsin fraction from the pyloric ceca of chinook salmon (Oncorhynchustshawytscha) CompBiochemPhysiolB, 143 :432-440
- nKlomklao, S.Benjakul, S.Visessanguan, W.Kishimura, H. Simpson, B.K., (2007). Trypsin from the pyloric caeca of bluefish (Pomatomussaltatrix) Comp BiochemPhysiol B, 148: 382-389
- nMalhotra, A. Coupland, J.N., (2004). The effect of surfactants on the solubility, zeta potential, and viscosity of soy protein isolates Food Hydrocolloid 18: 101-108
- nGhosh, S., (2008). Interaction of trypsin with sodium dodecyl sulfate in aqueous medium: A conformational view Colloids Surf, B 66: 178-186.